Standard Operating Procedure Approvals
The undersigned acknowledge that they have reviewed the Curation of Biomedical Data into ISA-TAB-Nano and caNanoLab SOP and agree with the information presented within this document. Changes to this Curation of Biomedical Data into ISA-TAB-Nano and caNanoLab SOP will be coordinated with and approved by the undersigned or their designated representatives.
Released by / Effective Date:
Written by
Name | Title | Date |
|---|---|---|
Michal Lijowski | Curator |
|
Approved by
Name | Title | Date |
|---|---|---|
Kanakadurga Addepalli | NCI Project Officer |
|
Standard Operating Procedure Revision History
Version Number | Implemented By | Revision Date | Description of Change |
|---|---|---|---|
0.5 | Michal Lijowski | Initial draft of "Nanotechnology Biomedical Data Curation Standard Operating Procedure" | |
0.6 | Michal Lijowski | Updated based on feedback | |
1.0 | Sharon Gaheen | Updated based on NCI feedback and re-formatted document in new template | |
2.0 | Michal Lijowski and Carolyn Klinger | Renamed as "Curation of Biomedical Data into ISA-TAB-Nano and caNanoLab Standard Operating Procedure," updated all procedures, and converted to NCI wiki. |
Purpose
This Standard Operating Procedure (SOP) explains how to curate biomedical data into ISA-TAB-Nano and caNanoLab.
References
The following table summarizes the documents referenced in this document.
Document Name | Description | Location |
|---|---|---|
ISA-TAB-Nano Information | This wiki contains information about ISA-TAB-Nano, which is an extension of ISA-TAB (Investigation/Study/Assay (ISA) tab-delimited (TAB) format), an existing specification developed by the ISA-community. | |
| Specification Documentation, ISA-TAB 1.0, Release Candidate 1 | This document contains guidelines on the format of ISA-TAB. | |
| caNanoLab User's Guide | This wiki contains procedures for using the caNanoLab data sharing portal. | |
caNanoLab Website | This site is the caNanoLab data sharing portal. | |
BioPortal Website | This website contains detailed descriptions of ontologies and forms associated with a publication or an investigation. |
Responsibilities
It is the responsibility of the Curation Subject Matter Expert (SME) to perform the steps summarized in Data Curation and detailed throughout this guide.
It is the responsibility of the NCI collaborator to provide a list of publications suggested for curation and contact investigators.
It is the responsibility of the Government Sponsor to supervise the technical and budgetary aspects.
It is the responsibility of the Project Officer to review the monthly report, if one is required.
It is the responsibility of the Leidos Technical Project Manager to coordinate technical and budgetary aspects.
Definitions
The following table provides definitions and explanations for terms and acronyms relevant to the content presented within this document.
Term | Definition |
|---|---|
ISA-TAB | Investigation Study Assay Tab delimited file format |
| ISA-TAB-Nano | ISA-TAB-Nano extends ISA-TAB, an existing specification developed by the ISA-community. Refer to the ISA-TAB Specification for guidelines on the format of ISA-TAB. The ISA-TAB-Nano wiki page provides additional information. |
NPO | Nanoparticle Ontology |
Curation Approach
Curation of biomedical information is accomplished by selecting relevant publications, extracting reported text and data, submitting extracted information to ISA-TAB-Nano and caNanoLab, and keeping track of performed activities.
caNanoLab (https://cananolab.cancer.gov) and ISA-TAB-Nano have differences in nomenclature, structure, and the way information is stored.
- In caNanoLab, all information regarding an individual material entity is stored in a single object called SAMPLE. This includes material composition, its functionalities, links between its constituents, its physicochemical, and in vitro, in vivo, and ex vivo properties derived from experiments reported in the publication. A term “Composition” annotates an object with SAMPLE, which contains information about material composition, its constituents, functionalities, and links between its constituents. A term “Characterization” annotates an object within SAMPLE, which contains information related to an individual assay performed to acquire data providing information about material properties.
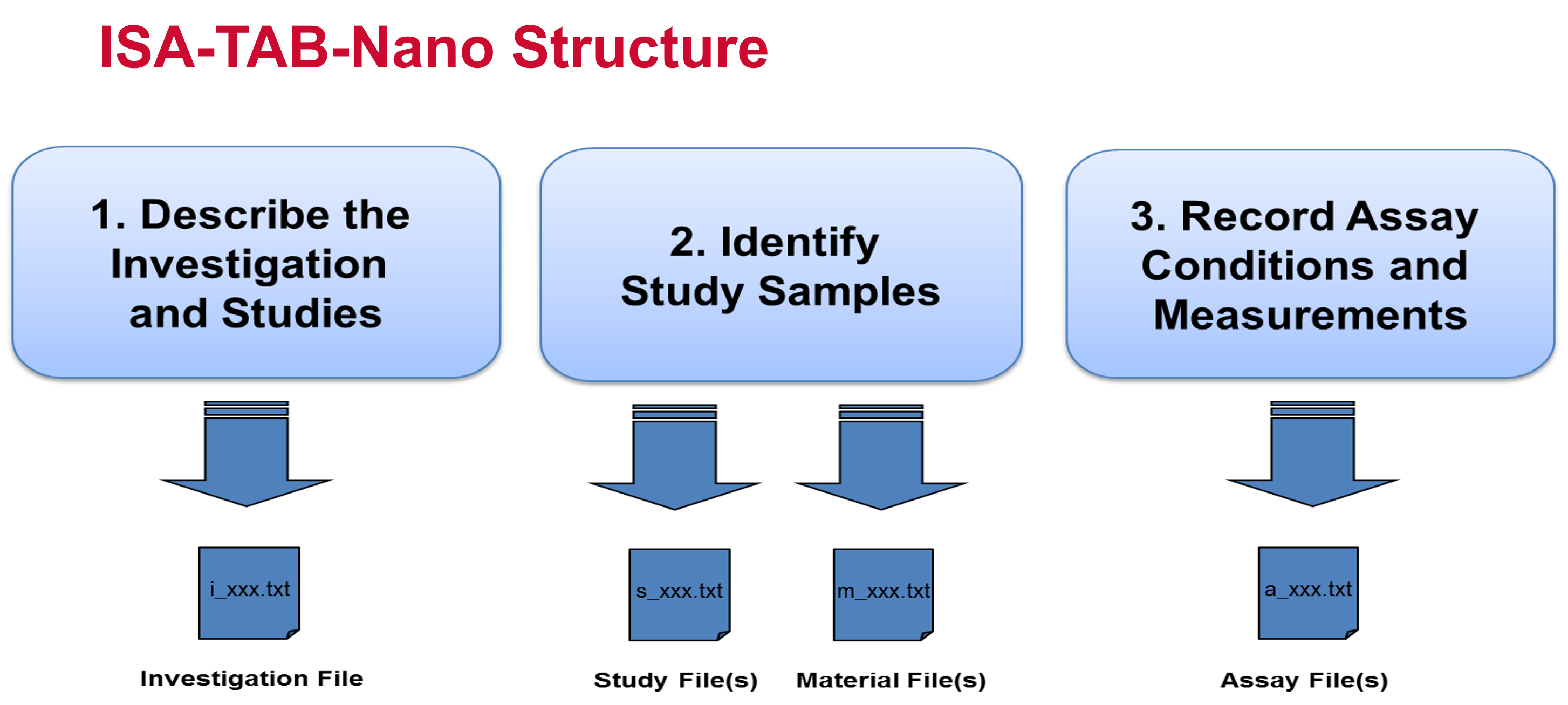
- In ISA-TAB-Nano (Figure 1), all information from a particular publication or an investigation is spread among Investigation, Study, Assay, and Material forms. The detailed description of each form, excluding the Material form, and their associations are provided in Specification documentation: release candidate 1, ISA-TAB 1.0 . A Material form corresponds to a “Composition” object in caNanoLab. Descriptive information stored in caNanoLab “Characterization” object components, like Design Description and Analysis and Conclusion, are stored in the Study Section of the Investigation form. Numerical data are stored in the Assay form. A Study form provides a link between the Investigation and Assay forms.
Figure 1. ISA-TAB-Nano Structure
Data Curation
Data curation is performed by following the steps below.
- The NCI collaborator provides a list of publications suggested for curation.
- The Curation SME evaluates publications on whether they are curatable, that is, comprise information relevant for curation in caNanoLab.
- The NCI collaborator contacts the investigator related to the selected publication to establish whether the investigator is willing to share numerical data and the additional information.
- The Curation SME extracts data from the publication.
- The Curation SME submits extracted data to caNanoLab and ISA-TAB-Nano forms. Information submitted to caNanoLab must be machine-readable, searchable, and comply with established standards.
- The Curation SME sends a request to the authors of a publication for additional data.
- The Curation SME submits additional data, such as numerical data used to generate figures and any missing information, to caNanoLab and ISA-TAB-Nano forms.
Primary Curation Steps
- Create a caNanoLabData folder on a system or server that gets backed up regularly. The caNanoLabData folder contains folders, named after the institution or collaboration, for example, USC_UV, which contain additional subfolders. These subfolders are named after first author of a publication, publication abbreviation (refer to Publication Abbreviations for suggested abbreviations to use), and publication year. For example, JCrecente-CampoJCR2019 contains an individual publication, which is a PDF file and any supplemental data associated with a publication, extracted data, and supplemental data provided by an author. If the data acquired for curation are not related to the publication, then name this subfolder after the institution, the person providing data, and the date of acquisition. The caNanoLabData folder contains all auxiliary files, such as a list each of cell lines, all curated publications, chemical compounds, new terms, and recently added, Bioportal terms.
- Create a subfolder in the caNanoLabData folder to store the publication and extracted data. A subfolder name comprises first author name, journal name, and year of publication. Create an additional subfolder within this subfolder to store ISA-TAB-Nano forms.
Data Extraction
The Curation SME performs the following data extraction steps.
- Establish a number of samples, which have different compositions or properties, and a number of characterizations using the information provided in the text, tables, figures, and figures' captions in curated publication and in supplementary information.
- Establish sample names using the following pattern: abbreviation(s) of institution names, name of the first author (without a middle name), custom abbreviation of the journal title, year of publication, and sample sequential number; for example, USC_UV-JCrecente-CampoJCR2019-01.
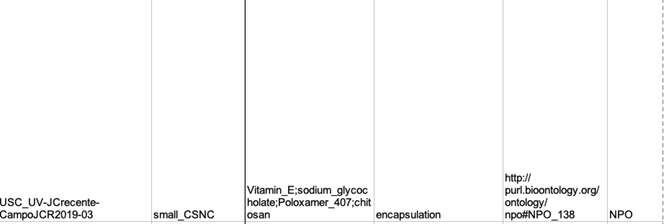
- Associate the samples with characterizations based on information provided in the text, tables, figures, and figure captions. This information is kept in a text file listing all samples and associated characterizations (Figure 2).
- Extract information on the composition, physicochemical, in vitro, and in vivo characterizations, numerical data for each individual sample into a corresponding text file. Remove references to figures and to publications. Rephrase active sentences to passive. For example “We synthesized the previously reported μMOF, Hf-DBA (DBA = 2,5-di(p-benzoato)aniline), and used it as a control.” replace with “The previously synthesized and reported μMOF, Hf-DBA (DBA = 2,5-di(p-benzoato)aniline), was used as a control”.
Establish definitions for new terms used in the publication, which are not in the caNanoLab glossary or Bioportal, but in other sources, like Wikipedia, and references therein, Encyclopedic Dictionary of Genetics, Genomics, and Proteomics. Record this definition or term in a designated text file, or if possible, enter into caNanoLab, for example, in targeting functionalized entity, a new target, such as a gene.
If the information provided by the publication, such as the name of an instrument or a chemical compound, does not agree with the information provided somewhere else, such as manufacturer catalog, retain the information provided by the publication for curation. Record the discrepancy for correspondence with authors in a file in the subfolder comprising all files comprising information to be submitted to caNanoLab and ISA-TAB-Nano that contains a request for the numerical data that were used to generate the figures.
Figure 2. Typical text showing associations between samples and characterizations
Data Submission
caNanoLab Data Submission
Submit the extracted information and reported numerical data into caNanoLab following the procedures in the caNanoLab User's Guide, which is accessible by selecting caNanoLab FAQ or Online Help buttons (Figure 2).
If you submit a new term in any field in caNanoLab, use all lowercase.
- Login into caNanoLab. If you do not have a caNanoLab account and want to save items in caNanoLab, contact caNanoLab-Support@ISB-CGC.org. You will be assigned a user role that affects what actions you can perform in caNanoLab and sent a login ID and password.
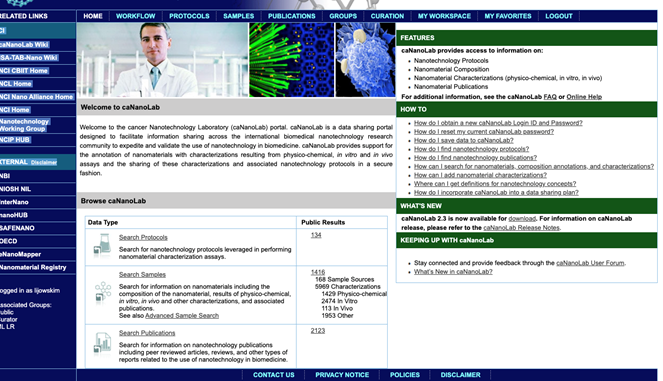
- Select either the SAMPLES tab on the top bar or the Submit Samples button (Figure 3).
Figure 3. caNanoLab home after login. Selecting tabs allows search samples, protocols, publications, and submission of samples, protocols, and publications.
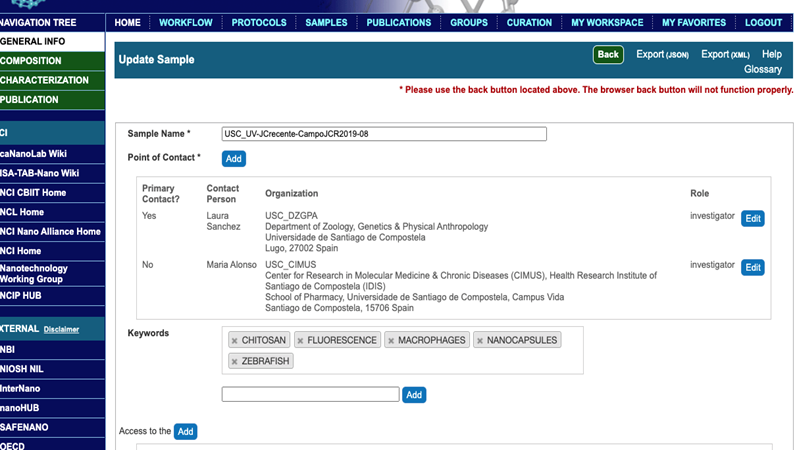
- In the General Info section, submit the sample name, contact information of the first author and the corresponding author(s) (full first name, first and middle initials, full last name, phone number, email address), name and custom-generated abbreviation for the institution(s) name(s), role (either manufacturer or investigator), addresses of corresponding authors, and keywords relevant to the publication. The first author would be assigned the data owner and the corresponding author is the primary point of contact (Figure 4).
Figure 4. A General Info window after submission of relevant data. Selecting Composition, Characterization, or Publication buttons on top left allows submission of sample composition, its characterizations, and the corresponding publication citation.
Composition Submission
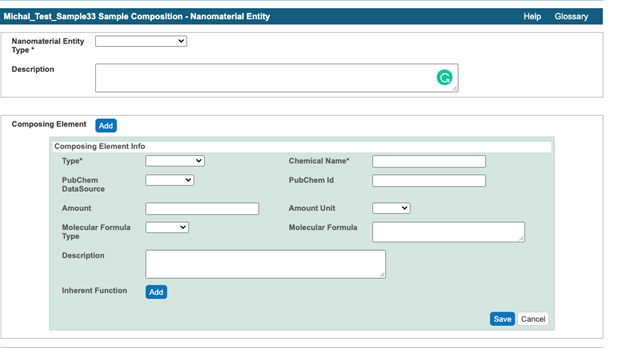
- Select the Composition button below General Info (Figure 5).
Figure 5. A window for submission of information about sample constituent - Select nanoparticle entity type from the Nanoparticle Entity Type drop-down menu and enter the particle description into the Description field.
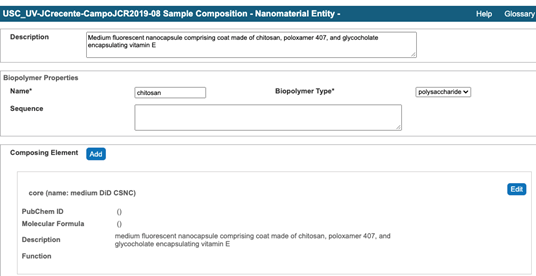
- Submit the sample composition into the Nanomaterial entity section. This includes the chemical name of the sample component, its type from a drop-down menu, its full name in the description field, PubChem Data Source from a drop-down menu, PubChem ID, and amount. The first Composing Element comprises information about a whole particle (Figure 6).
Figure 6. An example of the first composing element
- If any of the sample components has a function, e.g. targeting, the information is indicated in Inherent Function field (Figure 7). Select Function Type from the drop-down menu. Add Function Description and select Imaging Modality in the case of an Imaging Function (Figure 7, Figure 8).
Figure 7. A typical Composing Element window comprising an imaging inherent function
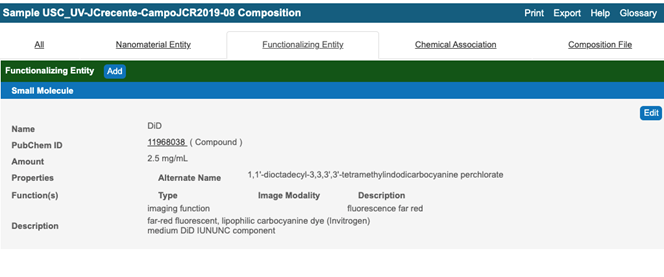
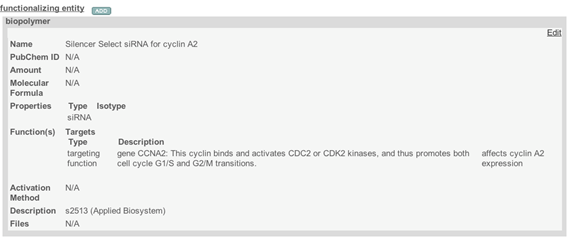
Figure 8. A typical window of Composing Element comprising a targeting imaging function - If a Composing Element having an Inherent Function is associated with another Composing Element, make a Composing Element with an Inherent Function as a Functionalized Entity. In most cases, select “small molecule” as the Functionalizing Entity Type, and submit additional information (Figure 9, Figure 10).
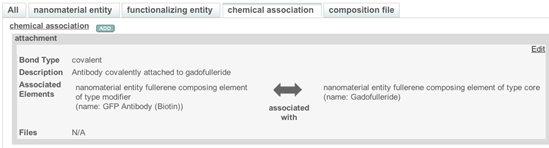
Select the Chemical Association tab, then click Add and select an Association Type from the drop-down menu. Add information about the association in the Description field. Select the Functionalized Element in the Element field on the left side and select the corresponding Composing Element on the right side in the Element field (Figure 11).
Figure 9. A typical window for a functionalized entity
Figure 10. A typical window for a targeting functionalized entity
Figure 11. A typical window for a chemical association
Characterization Submission
- Select the Characterization button (Figure 4).
- Select an appropriate Characterization Type, either physicochemical, in vitro , or in vivo from the drop-down menu (Figure 12).
- Select a Characterization Name from the drop-down menu. If a corresponding Characterization Name or Assay Type is not available, select either other_pc as physicochemical Characterization Name, other_vt for in vitro C haracterization Name, other_vv for in vivo C haracterization Name, or other_ex_vv for ex vivo Characterization Name.
- Select an assay type or, if an appropriate assay type is not available, enter a new assay type in the Assay Type field drop-down menu.
- Select Characterization Source from the drop-down menu.
- For in vitro characterization, enter a cell line, if a field for a cell line exists.
- Enter an assay description into the Design and Methods Description field.
- Enter technique(s) and instrument(s) used in the assay, either by selecting an existing technique and instrument, or by adding a new technique and instrument into the respective drop-down menus.
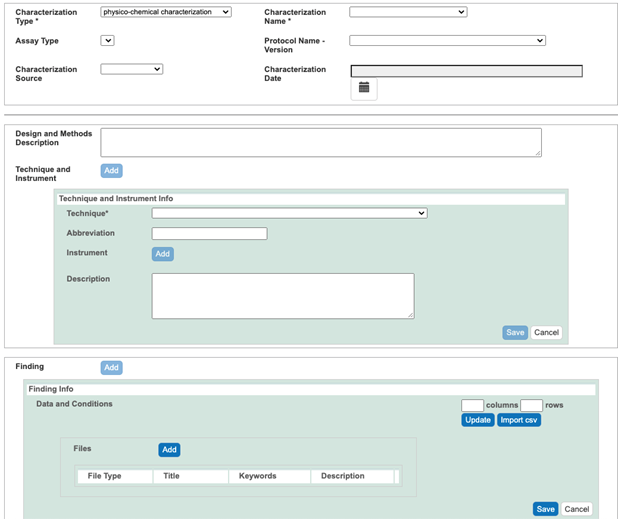
- If numerical data are available, click the Findings button. In the case of a small amount of data, enter the number of columns and rows required to accommodate these data, select the Update button, annotate columns, and enter data. In the case of a large amount of data, save it as a UTF-8 csv file (not just csv), click the Import csv button, and select the UTF-8 csv file to import (Figure 12). Regarding columns annotation, first select Column Type, either condition or datum. For example, if numerical data are provided as raw data, mean, uncertainty (standard deviation or standard error of the mean), and number of replications, then for each column containing either raw data, mean, standard deviation, and number of replicates, select values from the drop-down menus: Column Type datum, Column Name (e.g. diameter), Column Value Type (in the case of raw data, leave Column Value Type empty; if datum in the column is mean, then select mean), and Column Value Unit (Figure 13).
- Submit the description of the results from an assay into the Analysis and Conclusion field (Figure 13).
Figure 12. A Physicochemical Characterization window
Figure 13. A Physicochemical Characterization window with submitted data
Publication Submission
- Select the Publication tab (Figure 4).
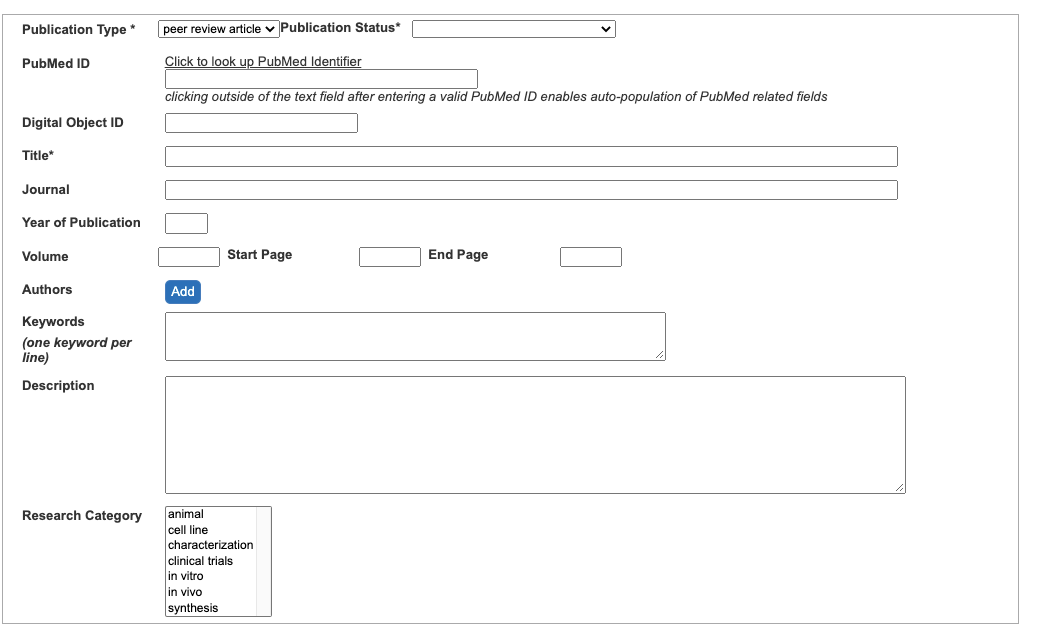
- Select a Publication Type for a drop-down menu.
- Select Publication Status from a drop-down menu.
- Enter PubMed ID and click outside PubMed ID field to obtain a citation for this publication (Figure 14).
Figure 14. An empty Publication window - If the publication does not exist in PubMed, then enter the publication DOI, its title, journal name, year of publication, volume, start and end pages, list of author names, keywords, abstract in Description field.
- Select Research Categories.
- Associate the publication citation with submitted samples as follows.
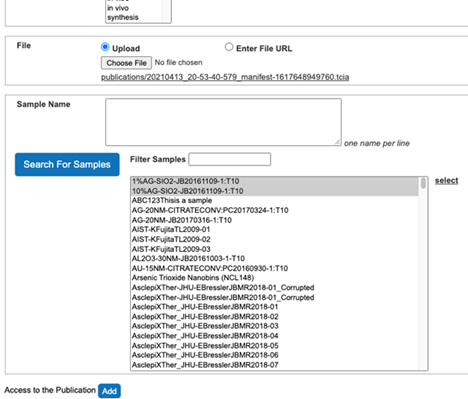
- Select Search For Samples button.
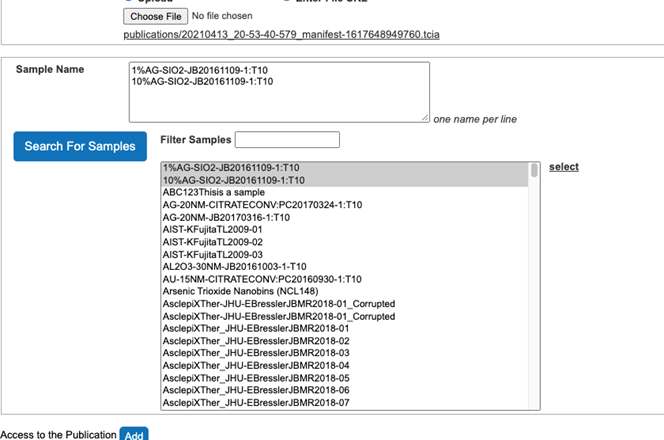
- Select Samples associated with the publication from the list of all samples (Figure 15).
Figure 15. Associate samples with the publication - Click the select button on the right side to associate Samples with the publication (Figure 16).
Figure 16. Samples are associated with the publication
- Set access to the publication citation as “public”.
- Review entries submitted into caNanoLab for consistency with information in the curated publication. Correct any issues.
- Generate Data Availability Metrics for each sample.
- Make all samples “public.”
Curation Queue
A Curation queue comprises Samples, Publications, and Protocols that users without Curator privileges submit. In order to make a Sample public, each section must have complete information. The sample Name must indicate the name of an institution that represents the origin of a sample. It should comprise other identifiable information that distinguishes it from other samples.
The General Info section should provide the name, postal address, and email address of an investigator or a person responsible for sample submission into caNanoLab. The sample must have its Composition and Characterization details submitted as mentioned in caNanoLab Data Submission, and be machine-readable, searchable, and comply with established standards.
Protocols
If a user submits a new version of a protocol to the caNanoLab curation queue, then the user must assign the old version of the protocol a version number, and assign the new version of the protocol a different version number.
ISA-TAB-Nano Data Submission
At the time this document was published, the curation in ISA-TAB-Nano follows the ISA-TAB-Nano 1.3 Release. More detailed information is provided in the ISA-TAB-Nano wiki. The filenames of the ISA-TAB-Nano forms consist of a prefix corresponding to a specific form, that is, i_ for an investigation form, s_ for a study form, a_ for an assay form, m_ for a material form, a custom abbreviation of institution(s) names, a name of the first author (first name abbreviation, full last name), a custom abbreviation of journal title and a year of publication; for example, i_USC_UV-JCrecente-CampoJCR2019. A suffix for Study indicates a study type e.g. physicochemical, in_vitro. A suffix for an Assay file and the type of Study it is related to includes a name of the assay; for example, size, zeta potential.
For example:
a_ USC_UV-JCrecente-CampoJCR2019-physicochemical-size-DLS
Investigation Form
The first lines of the Investigation form (Figure 17) are dedicated to names of ontologies from Bioportal (http://bioportal.bioontology.org). The information about ontologies is added while creating the ISA-TAB-Nano Investigation form and selecting the appropriate annotation from Bioportal for terms, which are entered into ISA-TAB-Nano forms. In case a term exists in multiple ontologies, select the most in depth annotation.
- Enter in the Term Source Name field an abbreviation of the ontology name.
- Enter the URL of the ontology in the Term Source File field.
- Enter the current version of the ontology or ontology release date in the Term Source Version field.
- Enter the full ontology name into the Term Source Description.
The most applicable ontologies are NanoParticle Ontology (NPO, https://bioportal.bioontology.org/ontologies/NPO ), NCI Thesaurus (NCIT, https://bioportal.bioontology.org/ontologies/NCIT), Eagle-I Research Resource Ontology (ERO, https://bioportal.bioontology.org/ontologies/ERO ), Ontology for Biomedical Investigations (OBI, https://bioportal.bioontology.org/ontologies/OBI ) Experimental Factor Ontology (EFO, https://bioportal.bioontology.org/ontologies/EFO ), Phenotypic Quality Ontology (PATO, https://bioportal.bioontology.org/ontologies/PATO ), and BioAssay Ontology (BAO, https://bioportal.bioontology.org/ontologies/BAO ). If it is necessary to annotate entries in Material, Study, and Assay files with terms from Ontologies which are not in Ontology Source Reference section than one should enter these Ontologies into this section.
ONTOLOGY SOURCE REFERENCE | |
|---|---|
Term Source Name | EFO |
Term Source File | |
Term Source Version | 3.29.0 |
Term Source Description | Experimental Factor Ontology |
Figure 17. An Ontology Source Reference section in an Investigation form
Enter the Investigation information into the Investigation form.
- Investigation Identifier, in most cases, comprising institution names, first author, journal title, and publication year
- Custom investigation title, e.g. a rephrased publication title
- Custom investigation description, e.g. an abbreviated abstract
- Custom investigation outcome
- PubMed ID
- Publication DOI
- Author list
- Publication title
- Publication status (published, submitted, in press, in preparation)
INVESTIGATION | |
|---|---|
Investigation Identifier | USC_UV-JCrecente0CampoJCR2019 |
Investigation Title | Dependence of interaction with macrophages and biodistribution in zebrafish on size and composition of polymeric nanocapsules |
Investigation Description | This work aimed to understand |
Investigation Submission Date | |
Investigation Public Release Date | |
Investigation Disease | |
Investigation Disease Term Accession Number | |
Investigation Disease Term Source REF | |
Investigation Outcome | The in vitro results showed that small nanocapsules interacted more efficiently with macrophages than their larger counterparts. Inulin nanocapsules were significantly less toxic than chitosan nanocapsules. Finally, following in vivo administration (intravenous/intramuscular) to zebrafish, small nanocapsules, regardless of their composition, disseminated considerably faster and further than their medium size counterparts. These results emphasize how small changes in the nanometric range can lead to a remarkably different interaction with the immune cells and biodistribution profile. |
Figure 18. A general information section of an Investigation form
Study Section of Investigation Form
Based on information obtained earlier and related to sample characterizations, identify studies and assays, which are common to a specific study.
Study Identifier | USC_UV-JCrecente0CampoJCR2019-physicochemical |
|---|---|
Study Title | size |
Study Description | The particle size and polydispersity index (PDI) were measured by a dynamic light scattering using a Zetasizer Nano S (Malvern) at 25 C with a detection angle 173 degrees and in distilled water. |
Study Submission Date | |
Study Public Release Date | |
Study Disease | |
Study Disease Term Accession Number | |
Study Disease Term Source REF | |
Study Outcome | |
Study File Name | s_USC_UV-JCrecente-CampoJCR2019-physicochemical.ods |
Figure 19. A Study Information subsection of a Study Section
Enter into a Study section of the Investigation form the following.
- Study Identifier, which can comprise the Investigation Identifier and study name, such as size
- Custom Study Title
- Custom Study Description – this may include a short description of all assays, which are included in the study, in case, when several assays are included in a single study.
- Study Disease, if is available, corresponding Term Accession Number from the Bioportal Ontology. (http://bioportal.bioontology.org) and Term Source REF, that is, the name of a corresponding ontology.
- Custom Study Outcome
- A Study Filename and its Description are entered after a corresponding Study form is created.
- Study Publication section is left blank, since there is no other publication related to this study besides a publication listed in the Investigation Publications section.
INVESTIGATION PUBLICATIONS | |
|---|---|
Investigation PubMed ID | 31306677 |
Investigation Publication DOI | 10.1016/j.jconrel.2019.07.011 |
Investigation Publication Author List | Jose Crecente-Campo;Jorge Guerra-Varela;Mercedes Peleteiro;Carlha Gutierrez-Lovera;Iago Fernandez-Marino; Andrea Dieguez-Docampo;Africa Gonzalez-Fernandez;Laura Sanchez;Maria Jose Alonso |
Investigation Publication Title | The size and composition of polymeric nanocapsules dictate their interaction with macrophages and biodistribution in zebrafish |
Investigation Publication Status | published |
Investigation Publication Status Term Accession Number | |
Investigation Publication Status Term Source REF | EFO |
Figure 20. A Study Publication section
Identify factors, that is, independent variables manipulated by the investigator with the intention to affect biological systems in a way that they can be measured by an assay. Enter them into Study Factor section (Figure 18). One factor per cell/column e.g. temperature, corresponding Accession Number and Term Source REF from Bioportal, Study Factor Type, its Accession Number, Term Source REF from Bioportal.
STUDY FACTORS | ||
|---|---|---|
Study Factor Name | nanoparticle sample | sample number |
Study Factor Type | nanoparticle sample | sample number |
Study Factor Type Term Accession Number | ||
Study Factor Type Term Source REF | NPO | MS |
Figure 21. A Study Factors section with two factors
Enter into the Study Assay section (Figure 22) the following: Study Assay Measurement Type, e.g. hydrodynamic size, corresponding Term Accession Number, Term Source REF, Study Assay Technology Type, for example, dynamic light scattering, corresponding Term Accession Number, and Term Source REF.
- Study Assay Technology Platform; that is, instrument name such as Zetasizer Nano ZS (Malvern), corresponding Term Accession Number, and Term Source REF.
- Study Assay Measurement Name, outputs from assay measurements; for example, hydrodynamic diameter, corresponding Term Accession Number, Term Source REF.
- Study Assay Filename, which is entered after creating the corresponding Assay form.
STUDY ASSAYS | |
|---|---|
Study Assay Measurement Type | hydrodynamic size |
Study Assay Measurement Term Accession Number | |
Study Assay Measurement Term Source REF | NPO |
Study Assay Technology Type | dynamic light scattering |
Study Assay Technology Type Term Accession Number | |
Study Assay Technology Type Term Source REF | NPO |
Study Assay Technology Platform | Zetasizer Nano S (Malvern) |
Study Assay Measurement Name | hydrodynamic diameter; polydispersity index |
Study Assay Measurement Name Term Accession Number | http://purl.bioontology.org/ontology/npo#NPO_1915;http://purl.bioontology.org/ontology/npo#NPO_1155 |
Study Assay Measurement Name Term Source REF | NPO;NPO |
Study Assay File Name | a_USC_UV-JCrecente-CampoJCR2019-physicochemical-size_DLS.ods |
Figure 22. A Study Assays section
If a protocol is provided (Figure 23), enter the following.
- Study Protocol Name
- Study Study Protocol Type and its corresponding Term Accession Number and Term Source REF from Bioportal
- Study Protocol Description
- Study Protocol Parameter Name, names of measurable quantities, which remain constant as part of assay, separated by semicolons
- Study Protocol Components Name, names of instruments, software, reagents etc., which are part of assay, separated by semicolon. Manufacturer names are entered next to the Component Name in parentheses.
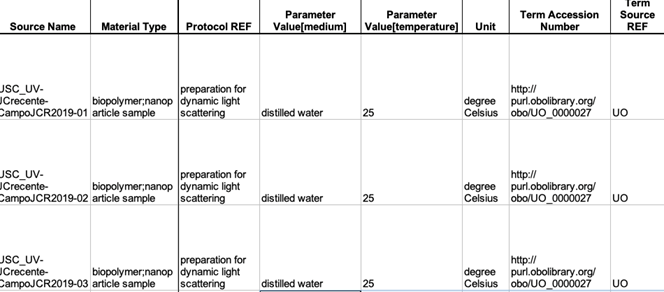
- Study Protocol Components Type, an instrument type, e.g. a flow cytometer, separated by semicolon, corresponding Term Accession Numbers from a Bioportal separated by semicolon into Term Accession Number field, Term Source REF, and Term Accession Number. In the empty columns to the right of protocols, enter the Study Protocol Name as preparation for the actual Protocol. Enter the Protocol Name in the Protocol REF field of the Study file (Figure 28).
STUDY PROTOCOLS | |
|---|---|
Study Protocol Name | dynamic light scattering |
Study Protocol Type | dynamic light scattering |
Study Protocol Type Term Accession Number | |
Study Protocol Type Term Source REF | NPO |
Study Protocol Description | |
Study Protocol URI | |
Study Protocol Version | |
Study Protocol Parameters Name | medium; temperature; detection angle |
Study Protocol Parameters Name Term Accession Number | |
Study Protocol Parameters Name Term Source REF | NPO;PATO; |
Study Protocol Components Name | distilled water;Zetasizer Nano S (Malvern) |
Study Protocol Components Type | medium; dynamic light scattering instrument |
Study Protocol Components Type Term Accession Number | |
Study Protocol Components Type Term Source REF | NPO;NPO |
Figure 23. A Study Protocols section
Figure 24. A Study Protocols section
Enter into Study Contacts section information from a contact person.
STUDY CONTACTS | |
|---|---|
Study Person Last Name | Doe |
Study Person First Name | Jane |
Study Person Mid Initials | JS |
Study Person Email | Jane.Doe@usc.es |
Study Person Phone | |
Study Person Fax | |
Study Person Address | Department of Zoology, Genetics & Physical Anthropology Universidade de Santiago de Compostela |
Study Person Affiliation | |
Study Person Roles | investigator |
Study Person Roles Term Accession Number | |
Study Person Roles Term Source REF | NCIT |
Figure 25. A Study Contacts section
Material Form
Create a number of Material forms corresponding to a number of identified samples. In addition to identified samples, which are submitted into caNanoLab, one can create Material forms for substances, like sucrose, saline, or drugs, which are used as control materials.
In the first line of Material form enter the information about a sample.
- Enter a first column Material Source Identifier, that is, a sample name as generated in section Data Extraction, for example, USC_UV-JCrecente-CampoJCR2019-04.
- Enter a material name, such as core/shell_iron//iron_oxide into the Material Name field.
- Enter the Manufacturer Lot Identifier, if one is available.
- In the Material Description field, enter a sample description.
- In the Material Synthesis field, enter a description of a sample synthesis.
- In the Material Design Rationale field, enter a description of the rationale to design a specific sample.
- Enter the Material Intended Application, its Term Accession Number, and the Term Source REF from Bioportal.
- Enter material type, e.g. nanoparticle sample, iron nanoparticle, into Material Type field. If several types are assigned to a sample, separate types by semicolon. Enter the corresponding Term Accession Number and Term Source REF, separated by a semicolon.
- In the Characteristics/Material Characteristic field, enter sample properties, for example, molecular weight.
- In the next lines, submit information about sample components, as above.
- In the Material Chemical Name field, enter the chemical name of a sample component, the corresponding Term Accession Number from Bioportal, and the Term Source REF. Check if the Ontology with this Term Accession Number is present in the Ontology Source Reference in the Investigation form (Figure 13). If it is not, then add this Ontology. Perform this process for all other forms and other Term Accession Numbers.
- In the Characteristics fields, enter additional information about this constituent of sample, such as amount, functionality, or molecular formula.
- If the amount of constituent is specified, enter unit into Unit field and corresponding Term Accession Number from Bioportal and the Term Source REF.
- If any two or more components of sample are linked, then in the very first line of Material form, enter Material Names of components, which are linked, into Material Part Name, and add information about a type of association into Material Linkage Type. Enter Term Accession Number corresponding to Material Linkage Type, and its Term Source REF (Figure 26). Repeat this step if the additional components are linked.
Figure 26. Part of a Material form showing a linkage subsection
Study and Assay Forms
The number of created Study forms should be equal to the number of Study sections in the Investigation form. In the case of imaging studies, create corresponding Study and Assay forms, when images are available without any restriction. Then enter the name of the file containing the image into the Image File field of a corresponding Assay form.
Study Form
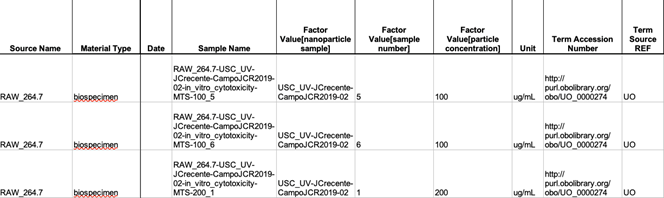
The number of fields in a Study form and which fields are in this form depends on the number of Study Factors, which are entered in the Investigation form for this particular Study. The common fields in the Study form are Source Name, Sample Name, Factor and Parameter Values (Figure 22), if both of the latter are specified in the Investigation form.
- In the Source Name field, enter either the Material Source Identifier in the case of a physicochemical Study; for example, USC_UV-JCrecente-CampoJCR2019-04, a cell line in the case of in vitro characterization e.g. H-1650, or an animal name, such as mice in the case of in vivo characterization.
- Enter the material type, either as a free-text description or as a term from Bioportal, its Term Accession Number, and Term Source REF.
- Enter into the Characteristics field cell type and cell line in the case of in vitro Study or animal type in case of in vivo.
- Enter the Factor Value, such as Material Source Identifier, in the case where an experiment involved only different samples, or entered multiple Factor values. In the Sample Name field, enter either the name of the nanomaterial sample, in the case where there are no Factors, or a name, which includes a combination of sample name, factors numerical values, cell name, and/or animal name. Number of lines in the Study form depends on the number of Factor Values and numerical Factor Values.
- Enter the protocol name from the corresponding Study Section in the Investigation form into the Protocol REF field.
- Enter the Study filename into the corresponding Study File Name field in the Investigation form.
Figure 27. Part of a Study form showing primary entries in this form for in vitro assay
Figure 28. Part of a Study form showing entries in this form for physicochemical assay
Assay Form
For each study, create a number of Assays forms corresponding to assays in the Study Assay subsection of the Study section in the Investigation form, in case corresponding numerical data are readily available. Enter the following into an Assay form (Figure 25).
- Enter the sample names created, while generating the corresponding Study form into the Sample Name fields.
- Enter the protocol name from the corresponding field in the Investigation form into the Protocol REF field.
- Enter the assay name into Assay Name fields.
- In the Measurement Value field, replace “measurement term” with appropriate term.
- Enter respective data into Measurement Value fields. Specify statistic, i.e. mean, standard deviation, and number of replicates in Statistic fields.
- If the numerical data possess units, enter units for mean and standard deviation into the Unit field, as well as the corresponding Term Accession Number and Term Source REF.
Sample Name | Protocol REF | Assay Name | Measurement Value[mean(hydrodynamic diameter)] | Unit | Term Accession Number | Term Source REF |
|---|---|---|---|---|---|---|
USC_UV-JCrecente-CampoJCR2019-01-physicochem-DLS-size | dynamic light scattering | hydrodynamic diameter measurement | 69 | nm | UO | |
USC_UV-JCrecente-CampoJCR2019-02-physicochem-DLS-size | dynamic light scattering | hydrodynamic diameter measurement | 246 | nm | UO |
Final Steps
- Complete the Investigation form by entering Material File Names, Material Source Name, Study File Names, and Assay File Names.
- Create a ISA-TAB-Nano_csv folder on your computer.
Convert all files to csv format. This step can be performed in macOS and Linux platforms using the unoconv script . Run this script from the ISA-TAB-Nano folder in a terminal window.
unoconv -e FilterOptions=9/32,,9 -f csv -o ../ISA-TAB-Nano_csv*.xlsx
Use *.xlsx if ISA-TAB-Nano forms were created using Excel.- In the Investigation form, replace all extensions, such as xlsx, with csv using a text editor or a script.
- Compress all forms and associated information into a single compressed file, and post the file to the caNanoLab Data Curation Project Wiki under ISA-TAB-Nano Curated Examples, including a citation of the publication.
- Update the caNanoLab/ISA-TAB-Nano curation status in the caNanoLab Data Curation Project Status file.
- After completing caNanoLab curation, provide the NCI collaborator with an explicit list and description of the data needed from investigators to complete the curation task.
- In the NCI JIRA tracker, log issues regarding defects encountered in caNanoLab and requests for improvements.
Appendix A
Publication Abbreviations
The abbreviations below are suggested for use during caNanoLab curation.
| Journal Title | Custom Abbreviation |
|---|---|
| AAPS J | AAPSJ |
| ACS Nano | ACSNano |
| Adv Func Mater | AFM |
| Adv Mater | AM |
| Advanced Healthcare Materials | AHM |
| Anal Chem | AC |
| Angew Chem Int Ed Engl | ACIEE |
| Anticancer Drugs | AD |
| Arterioscler Thromb Vasc Biol | ATVB |
| Bioconjug Chem | BC |
| Biomacromolecules | BM |
| Biomaterials | Bmat |
| Cancer Lett | CL |
| Cancer Res | CR |
| Chem Commun | CCR |
| Chem Res Toxicol | CRT |
| ChemBioChem | CBC |
| Clin Cancer Res | CCR |
| Environ Health Perspect | EHP |
| Environ Mol Mutagen | EMM |
| Eur J Pharm Sci | EJPS |
| Faseb J | FJ |
| IEEE Ultrasonics Symposium | IEEEUS |
| Int J Cancer | IJC |
| Int J Nanomedicine | IJN |
| Int J Pharm | IJP |
| Integr Biol | IR |
| Invest Radiol | IR |
| J Agric Food Chem | JAFC |
| J Am Chem Soc | JACS |
| J Clin Invest | JCI |
| J Control Release | JCR |
| J Drug Target | JDT |
| J Mater Chem | JMC |
| J Nanobiotechnology | JNBT |
| J Nucl Med | JNM |
| J Pharm Sci | JPS |
| Langmuir | LGMR |
| Magn Reson Med | MRM |
| Mater Sci Eng C | MSEC |
| Mol Pharm | MP |
| Mol Ther | MT |
| Nano Lett | NL |
| Nanomedicine (London) | Nanomed |
| Nanomedicine: Nanotechnology, Biology and Medicine | NNBM |
| Nanoscale | Nanoscale |
| Nanotechnology | NT |
| Nanotoxicology | NTX |
| Nature | Nature |
| Nature Medicine | NatMed |
| Nature Nanotechnol | NatNano |
| Nature Biotechnol | NatBiotech |
| Nature Commun | NatComm |
| Nature Materials | NatMat |
| Nucleic Acid Res | NAR |
| Pharm Res | PR |
| Photochem Photobiol | PhPh |
| Proc Natl Acad Sci U S A | PNAS |
| Radiology | Radiol |
| Scanning | Sc |
| Sci Transl Med | STM |
| Science | Science |
| Scientific Reports | SR |
| Small | Small |
| Toxicol Appl Pharmacol | TAP |
| Toxicol Lett | TL |
| Toxicol Sc | TS |
| Ultrasound Med Biol | UMB |