|
Page History
...
This document is a section of the Administration Guide.
Vocabulary Administration Overview
...
Administrative Program | Description | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
ActivateScheme | Activates a coding scheme based on unique URN and version.
| |||||||||
ClearOrphanedResources | Clean up orphaned resources - databases and indexes.
| |||||||||
DeactivateScheme | Deactivates a coding scheme based on unique URN and version.
| |||||||||
ExportLgXML | Exports content from the repository to a file in the LexGrid canonical XML format.
| |||||||||
ExportOBO | Exports content from the repository to a file in the Open Biomedical Ontologies (OBO) format.
| |||||||||
ExportOWL | Exports content from the repository to a file in OWL format.
| |||||||||
ListExtensions | List registered extensions to the LexEVS runtime environment.
| |||||||||
ListSchemes | List all currently registered vocabularies.
| |||||||||
LoadLgXML | Loads a vocabulary file, provided in LexGrid canonical xml format.
| |||||||||
LoadNCIHistory | Imports NCI History data to the LexEVS repository.
| |||||||||
LoadNCIMeta | Loads the NCI MetaThesaurus, provided as a collection of RRF files.
| |||||||||
LoadNCIThesOWL | Loads an OWL file containing a version of the NCI Thesaurus ...
| |||||||||
LoadOBO | Loads a file specified in the Open Biomedical Ontologies (OBO) format.
| |||||||||
LoadOWL | Loads an OWL file.
Options:
| |||||||||
LoadUMLSDatabase | Loads UMLS content, provided as a collection of RRF files in a single directory. Files may comprise the entire UMLS distribution or pruned via the MetamorphoSys tool. A complete list of source vocabularies is available online at http://www.nlm.nih.gov/research/umls/metaa1.html.
| |||||||||
LoadUMLSFiles | Loads UMLS content, provided as a collection of RRF files in a single directory. Files may comprise the entire UMLS distribution or pruned via the MetamorphoSys tool. A complete list of source vocabularies is available online at http://www.nlm.nih.gov/research/umls/metaa1.html.
| |||||||||
LoadUMLSSemnet | Loads the UMLS Semantic Network, provided as a collection of files in a single directory. The following files are expected to be provided from the National Library of Medicine (NLM) distribution:
| |||||||||
LoadFMA | Imports from an FMA database to a LexEVS repository. Requires that the pprj file be configured with a database URN, username, password for an FMA MySQL based database. The FMA.pprj file and MySQL dump file are available at http://sig.biostr.washington.edu/projects/fm/ upon registration.
| |||||||||
LoadHL7RIM | Converts an HL7 RIM MS Access database to a LexGrid database
| |||||||||
LoadMetaData | Loads optional XML-based metadata to be associated with an existing coding scheme.
| |||||||||
RebuildIndex | Rebuilds indexes associated with the specified coding scheme.
| |||||||||
RemoveIndex | Clears an optional named index associated with the specified coding scheme.
Options:
| |||||||||
RemoveScheme | Removes a coding scheme based on unique URN and version.
| |||||||||
TagScheme | Associates a tag ID (e.g. 'PRODUCTION' or 'TEST') with a coding scheme URN and version.
| |||||||||
TestRunner | Located in {LEXBIG_DIRECTORY}/test. Executes a suite of tests for the LexEVS installation.
Options:
| |||||||||
| | TransferScheme | Tool to help gather information necessary to transfer data from one SQL server to another.
|
Installing Sample Vocabularies
This LexEVS installation provides the UMLS Semantic Net and a sampling of the NCI Thesaurus content (sample.owl) that can be loaded into the database.
- In a Command Prompt window, enter the following to go to the example programs:
Code Block cd {LEXBIG_DIRECTORY}/examples - To load the example vocabularies, run the appropriate LoadSampleData script (
...
- LoadSampleData.
...
- bat for Windows;
...
- LoadSampleData.
...
- sh for Linux).
| Info | ||
|---|---|---|
| ||
Vocabularies should not be loaded until configuration of the LexEVS runtime and database server are complete. |
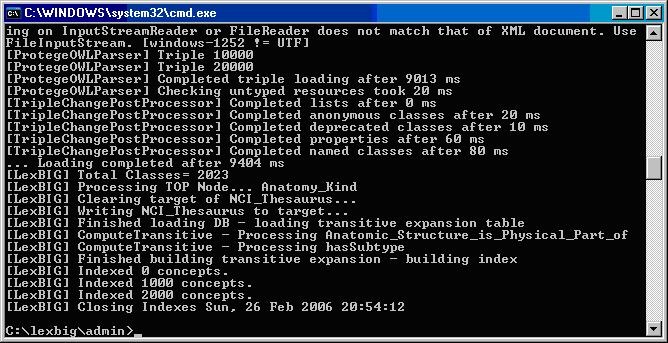
The following figure displays the successful load of the sample vocabulary file.
Running the Sample Query Programs
A set of sample programs are provided in the {LEXBIG_DIRECTORY}/examples directory. To run the sample query programs successfully a vocabulary must have been loaded.
...
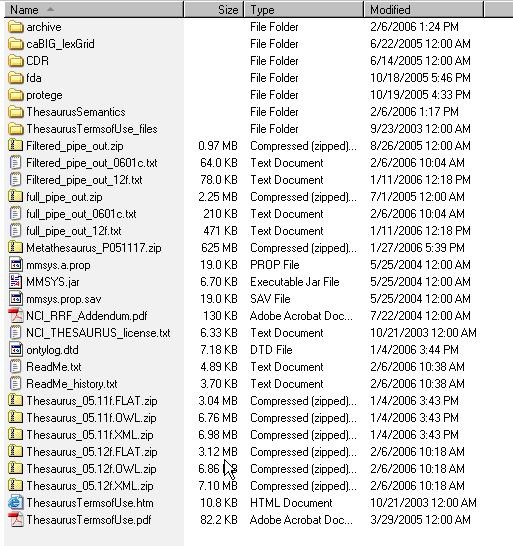
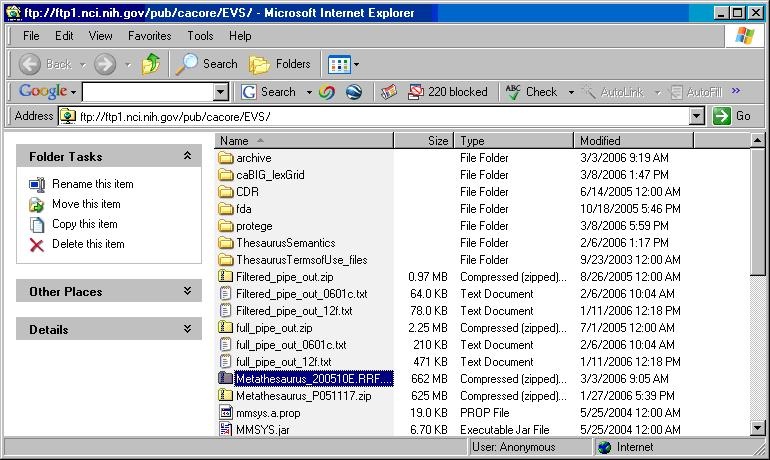
- Using a web or ftp client go to URL: ftp://ftp1.nci.nih.gov/pub/cacore/EVS/

- Select the version of NCI Thesaurus OWL you wish to download. Save the file to a directory on your machine.
- Extract the OWL file from the zip download and save in a directory on your machine. This directory will be referred to as NCI_THESAURUS_DIRECTORY
- Using the LexEVS utilities load the NCI Thesaurus
For Windows installation use the following command:Code Block cd {LexBIG_DIRECTORY}/admin
For Linux installation use the following command:Code Block LoadNCIThesOWL.bat -nf -in "file:///{NCI_THESAURUS_DIRECTORY}/Thesaurus_05.12f.owl"Code Block LoadNCIThesOWL.sh -nf -in "file:///{NCI_THESAURUS_DIRECTORY}/Thesaurus_05.12f.owl"Info title Note This step will require about three hours on a Pentium 3.0 Ghz machine. The total time to load NCI Thesaurus will vary depending on machine, memory, and disk speed.
...
| Code Block |
|---|
... [LexBIG] Processing TOP Node... Retired_Kind [LexBIG] Clearing target of NCI_Thesaurus... [LexBIG] Writing NCI_Thesaurus to target... [LexBIG] Finished loading DB - loading transitive expansion table [LexBIG] ComputeTransitive - Processing Anatomic_Structure_Has_Location [LexBIG] ComputeTransitive - Processing Anatomic_Structure_is_Physical_Part_of [LexBIG] ComputeTransitive - Processing Biological_Process_Has_Initiator_Process [LexBIG] ComputeTransitive - Processing Biological_Process_Has_Result_Biological_Process [LexBIG] ComputeTransitive - Processing Biological_Process_Is_Part_of_Process [LexBIG] ComputeTransitive - Processing Conceptual_Part_Of [LexBIG] ComputeTransitive - Processing Disease_Excludes_Finding [LexBIG] ComputeTransitive - Processing Disease_Has_Associated_Disease [LexBIG] ComputeTransitive - Processing Disease_Has_Finding [LexBIG] ComputeTransitive - Processing Disease_May_Have_Associated_Disease [LexBIG] ComputeTransitive - Processing Disease_May_Have_Finding [LexBIG] ComputeTransitive - Processing Gene_Product_Has_Biochemical_Function [LexBIG] ComputeTransitive - Processing Gene_Product_Has_Chemical_Classification [LexBIG] ComputeTransitive - Processing Gene_Product_is_Physical_Part_of [LexBIG] ComputeTransitive - Processing hasSubtype [LexBIG] Finished building transitive expansion - building index [LexBIG] Getting a results from sql (a page if using mysql) [LexBIG] Indexed 0 concepts. [LexBIG] Indexed 5000 concepts. [LexBIG] Indexed 10000 concepts. [LexBIG] Indexed 15000 concepts. [LexBIG] Indexed 20000 concepts. [LexBIG] Indexed 25000 concepts. [LexBIG] Indexed 30000 concepts. [LexBIG] Indexed 35000 concepts. [LexBIG] Indexed 40000 concepts. [LexBIG] Indexed 45000 concepts. [LexBIG] Indexed 46000 concepts. [LexBIG] Getting a results from sql (a page if using mysql) [LexBIG] Closing Indexes Mon, 27 Feb 2006 01:44:22 [LexBIG] Finished indexing |
NCI Metathesaurus
...
Vocabulary
This section describes the steps to download and install a full version of the NCI Metathesaurus for the LexEVS Service.
- Using a web or ftp client go to URL: ftp://ftp1.nci.nih.gov/pub/cacore/EVS/

- Select the version of NCI Metathesaurus RRF you wish to download. Save the file to a directory on your machine.
- Extract the RRF files from the zip download and save in a directory on your machine. This directory will be referred to as NCI_METATHESAURUS_DIRECTORY.
Info title Note RELEASE_INFO.RRF is required to be present for the load utility to work.
- Using the LexEVS utilities load the NCI Thesaurus
For Windows installation use the following command:Code Block cd {LexBIG_DIRECTORY}/admin
For Linux installation use the following command:Code Block LoadNCIMeta.bat -nf -in "file:///{NCI_METATHESAURUS_DIRECTORY}/"Code Block LoadNCIMeta.sh -nf -in "file:///{NCI_THESAURUS_DIRECTORY}/"
...
- Using a web or ftp client go to URL: ftp://ftp1.nci.nih.gov/pub/cacore/EVS/

- Select the version of NCI History you wish to download. Save the file to a directory on your machine. Select the VersionFile download to the same directory as the history file.
- Extract the History files from the zip download and save in a directory on your machine. This directory will be referred to as NCI_HISTORY_DIRECTORY
- Using the LexEVS utilities load the NCI Thesaurus:
For Windows installation use the following command:Code Block cd {LexBIG_DIRECTORY}/admin_*</tt>
For Linux installation use the following command:Code Block LoadNCIHistory.bat -nf -in "file:///{NCI_HISTORY_DIRECTORY}" -vf "file:///NCI_HISTORY_DIRECTORY}/VersionFile"Code Block LoadNCIHistory.sh -nf -in "file:///{NCI_HISTORY_DIRECTORY}" -vf "file:///NCI_HISTORY_DIRECTORY}/VersionFile"
...
- Change directory to LexEVS administration directory and enter:
Code Block cd {LEXBIG_DIRECTORY}/admin_* - Use the TagScheme to tag a coding system and version with a local tag name (e.g. PRODUCTION). This tag name can be used via LexEVS API for query restriction. Example:
|Code Block TagScheme -u "urn:oid:2.16.840.1.113883.3.26.1.1" -v "05.12f" -t "PRODUCTION"
| Scrollbar | ||
|---|---|---|
|