cloak.toggle.type = text cloak.toggle.open=[show] cloak.toggle.close=[hide] |
|
Topic:
Release:
Date entered:
Please leave your comment in the caArray End User Forum.
<strong>Applicable Releases:</strong> caArray 2.X
<strong>Date entered:</strong> 02/12/2009
</span>
questionDiv
question}}Question: *What are the MAGE-TAB Files? * {{questionEnd
questionDivEnd
<noinclude>
|
genomic locations. Columns represent samples or experimental conditions. | Yes | No. Directly Import |
|
The term of "MAGE-TAB Files" (See an example)has been used to refer not only MAGE-TAB formatted files as summarized in table 1, but also files that are supported by MAGE-TAB parser mentioned in last section. To be more specific, MAGE-TAB files also include the third party's raw array data files, derived data files and array design files from array providers, as shown in the figure below. |
In summary, MAGE-TAB files refer to each other. Together they represent the complete experiment.
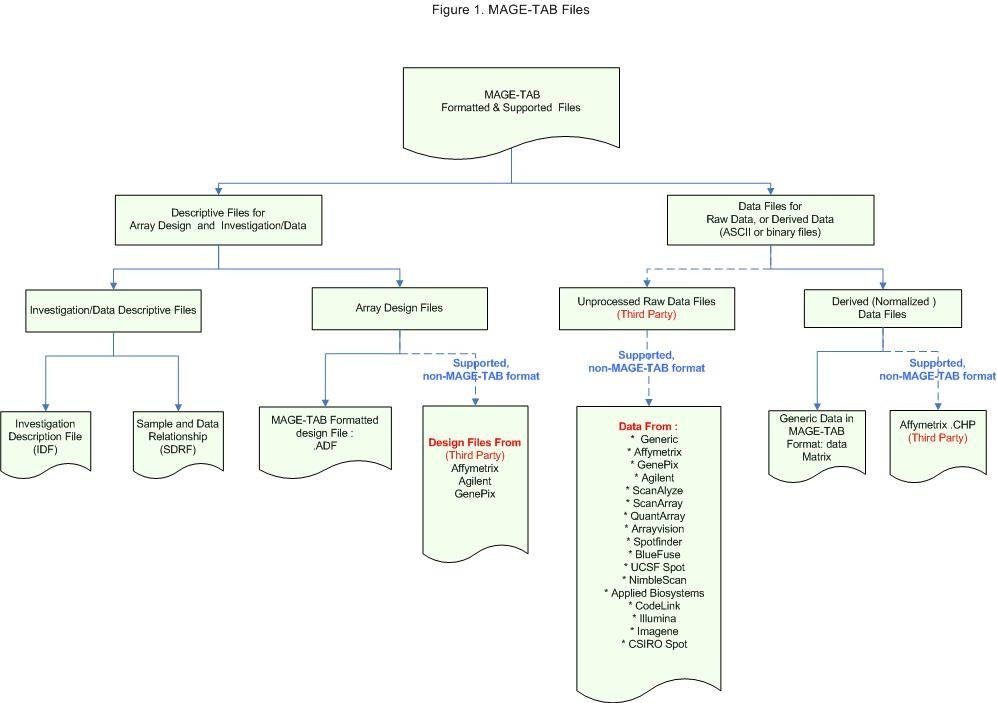
MAGE-TAB specification can be found at: MGED homepage.
To get start, you may generate a MAGE-TAB template file from EMBL-EBI's MAGE TAB site, or create your own IDF & SDRF files based on the MAGE-TAB documentation.
MAGE-TAB trainings and demos for the caArray users are currently under the development. We will add the links here once they become available.
|
Read more about When to use MAGE-TAB annotation files in caArray, Click Here |
|
Have a comment? Please leave your comment in caArray End User Forum |
</html>