This chapter describes search results that caIntegrator returns after queries.
Topics in this chapter include the following:
After you launch a search of a caIntegrator study, the system automatically opens the Query Results tab showing the results of your search. If you have not configured the column and sort display parameters before launching the search, by default the tab shows only the subject identifiers and a column that allows you to select each row of the data subset.
To display and/or sort additional data, you must return to the Results Type Tab and/or Sorting Tab to set display parameters, then re-run the search. The new search results will display the additional information, with the columns and data sorted as you specified.
caIntegrator paginates search results into pages of configurable size (default 20) with standard paginated navigation controls. To sort columns by ascending or descending parameters for on any displayed field, click on the underlined column header.
The query results that can display depend upon the criteria you established for the search. Follow the links below for more information about the category of data you searched.
You can download search results as a CSV file. The file contains the annotations, columns and data sort configurations you specified in the search query. See #Exporting Data.
See #Subject Annotation and Imaging Data, #Gene Expression Data, and #Expanding Imaging Data Results.
If you run the search before configuring column and sort display parameters, only the \[subject\] ID that meet the criteria and a column allowing you to select each row appear on the table, as shown in the following figure. !imaging subj ID only75.png|vspace=4, alt="Query Results page"! |
You can add details for one or more subjects by configuring them on the Results Type tab. Annotations listed there are the column headers in the CSV file(s) that were uploaded to the study. For information about using the Results Type tab, see Results Type Tab.
If after defining gene expression criteria on the Criteria tab, you select the Gene Expression result type on the Results Type tab, genomic data search results display in a gene expression data matrix. Because the data was downloaded from caArray, the data permissions granted there still apply. In other words, if you have been given access to the data in caArray, you can see it in caIntegrator.
You can select on the Results Type tab a preferred orientation for displaying the results: genes in rows and subjects in columns, or genes in columns and subjects in rows.
For Gene criteria, the cells display the median gene expression value for each gene. Next to each gene symbol, caIntegrator displays an icon (![]() ) which you can click to open the Cancer Genome Anatomy Project (CGAP) showing data for the gene. Icons are identified in the following figure.
) which you can click to open the Cancer Genome Anatomy Project (CGAP) showing data for the gene. Icons are identified in the following figure.
If you have selected Gene Expression on the Results Type tab, then the column headers are a clickable label which sorts the entire table on that column. If you selected Reporter ID on the Results Type tab, the Reporter ID is clickable (and the gene is not clickable).
For fold-change criteria, the cells display the normalized signal-based value for a given reporter for a given sample. In the results matrix, caIntegrator highlights matrix values for fold change results that meet fold change criteria. Red represents upregulated values and blue indicates downregulated values. The following two figures display gene name search results with gene reporter type display in the first and reporter ID reporter type display in the second. Note the left hand column in each example.
You can save genes identified in the search results as a gene list. For more information, see #Creating a Gene or Subject List.
After defining copy number criteria on the Criteria tab and running a copy number query, you should select the Copy Number result type on the Results Type Tab, and rerun the query. Copy number data search results display in a data matrix containing samples vs. genomic regions.
DNAcopy ouput values can be negative. If the test and the reference genomic samples both have two copies of a chromosomal region, the ratio of test/reference is '1', and the log2(1) = 0. That is, if there is no change in the chromosomal structure, then the value is 0. If there are more copies in the test sample (amplification of the chromosomal segment), the ratio of test to reference is greater than 1, and the log2(test/reference) is greater than 0. For example, if the test sample has 6, the ratio or test/reference is 6/2 = 3; log2(3) = 1.58. In a deletion, the test is less than the reference, for example 1. The DNAcopy output value would be log2(1/2) = log2(0.5) = -1.0. Values below -0.6 are often considered a deletion.
From any page in caIntegrator that shows such a group, you can save a list of genes or subjects so you can use it for searches or analyses. This functionality can also be used where a gene or subject list was created outside of caIntegrator, for example, a list of subjects with validated mutation such as from TCGA projects, or a list of subjects with high EGFR expression or any lists of subjects with genomic or clinical characteristics determined with other tools.
To create a list, follow these steps:
Click *Create List* at the bottom of the page. caIntegrator now opens the Edit \[Subject or Gene\] List page which shows the name and symbols of the newest gene list, shown in the following figure. !edit gene list80.png|vspace=4, border=1, alt="The Edit Gene List for reviewing, editing the name or deleting a gene list. The Edit Subject List page is comparable."! |
See #Editing a Gene or Subject List for information about the edit feature.
When you perform a GISTIC analysis, caIntegrator automatically saves the retrieved genes in the Saved Copy Number analysis in the left sidebar. For a query or plot analysis, they also appear in the Gene Picker dialog box described in #Choosing Genes. |
To view a gene list or subject list in caIntegrator, under Study Data in the left sidebar, click Saved Lists > Global Lists, or My Lists. Select the list/analysis you want to open. The system displays gene or subject lists that have been saved for the open study.
You can initiate the following functions on this page:
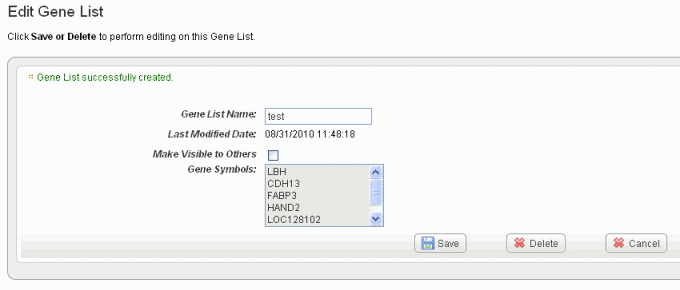
In the Edit \[List Type\] dialog box, you can perform the following tasks: |
To rename the list in the *\[List Type\] List Name* text box, enter the new list name. |
Once a list is created, you cannot edit the list contents.
Once you have run a query for gene expression, or for copy number, you can view results in the Integrative Genomics Viewer (IGV).
The IGV is a high-performance visualization tool for interactive exploration of large, integrated datasets. It supports a wide variety of data types including sequence alignments, microarrays, and genomic annotations.
For more information about the Integrative Genomics Viewer or to connect independently to the IGV home page, see Integrative Genomics Viewer login. You may also want to refer to the IGV User Guide. The IGV viewer and the NCI Heat Map viewer both require you to install a version of Java containing Java Web Start. For more information, see #Java for IGV and Heat Map Viewewr. |
There are two ways to integrate caIntegrator with the IGV. To configure the connection to IGV, follow one of these methods.
IGV.jnlp opens, displaying the dataset in the computer screen. An example displays in the following figure.Once you have run a query for gene expression, or for copy number, you can view results in the Heat Map Viewer (HMV).
For more information about the Heat Map Viewer or to connect independently to the HMV home page, see Heat Map Viewer documentation. For HMV documentation, see https://cgwb.nci.nih.gov/goldenPath/heatmap/documentation/index.html. The IGV viewer and the NCI Heat Map viewer both require you to install a version of Java containing Java Web Start. For more information, see #Java for IGV and Heat Map Viewer.. |
There are two ways to integrate caIntegrator with the Heat Map Viewer. To configure the connection, follow one of these methods.
For interpretation of the results and using HMV features, see the help files opened from HMV. |
To use the IGV and the NCI Heat Map viewer, described in #Viewing Data with the Integrative Genomics Viewer and #Viewing Data with Heat Map Viewer, you must install a version of Java containing Java Web Start. You must install recent versions of the Java Development Kit (JDK 1.5.0 aka JDK 5.0 or newer) or Java Runtime Environment (JRE 1.5.0 aka JRE 5.0 or newer). The easiest option is to install JRE 5.0. For more information, see http://www.java.com/en/download/faq/java_webstart.xml.
Without Java Web Start, when you click Launch Integrative Genomics Viewer or Launch Heat Map Viewer, a dialog box displays in your browser giving you the option to save or open with igv.jnlp (IGV) or retrieveFile.jnlp (HMV). Clicking the Open option starts the Java Web Start Launcher (default), installing the Java app so that you can view the files.
The first time you launch the IGV or HMV with Java properly installed, regardless of browser type, a warning may appear: the "the digital signature cannot be verified". Click Run to proceed with opening the viewer. |
In reviewing imaging search results, it is important to understand the hierarchy of submissions in NBIA. For more information, see #Relationship of Subject to Study to Series to Images.
If you run a search before configuring column and sort display parameters, only the Subject Identifiers for the patients/images that meet the criteria and a column containing one check box per row display by default. An example displays in the following figure.
If your annotation choice on the Results Type page identifies annotations such as tumor size or tumor location, the search results display image series subsets that have those annotations, or any annotations you check on the Results Type page. The check boxes work in conjunction with buttons at the bottom of the results page, shown in the following figure. By expanding display parameters, you can view complete details for image search results.
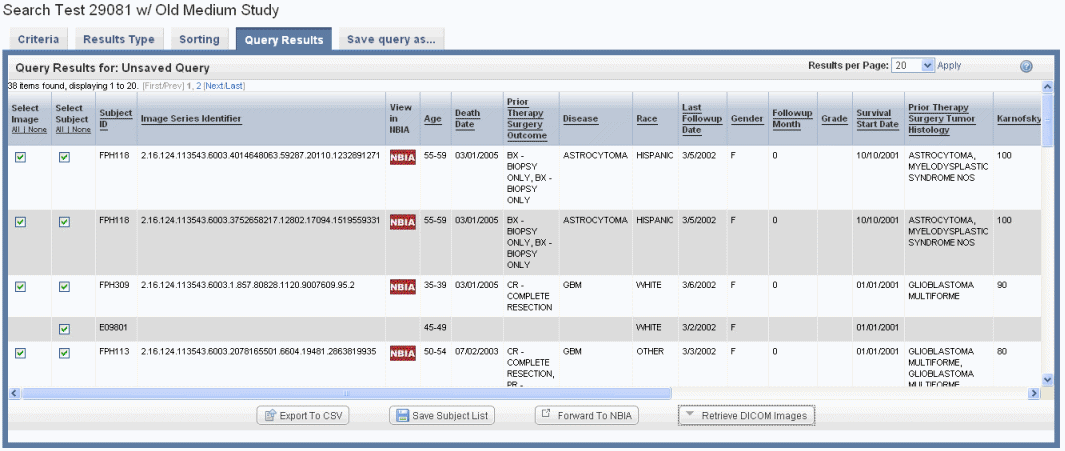
You can add more details for images by configuring image annotations on the Results Type tab. Annotations listed there are the column headers in the image series CSV file(s) that were uploaded to the study. Examples of image details include the following:
You can set display parameters for the results on the Columns and Sorting tabs. For more information, see #Results Type Tab.
See also #caIntegrator and NBIA, [#Retrieving DICOM Images and #Example of Retrieving Images.
Images can be accessed in NBIA if you see buttons on the Search Results page. See the Imaging Note in #Results Type Tab. You can click links on the Search Results tab to view or download image data.
34. Retrieving DICOM Images
On the caIntegrator imaging data Search Results page, you can click the Retrieve DICOM Images button which is linked to results you have selected by row. caIntegrator retrieves the corresponding image(s) from NBIA through the grid. NBIA organizes the download file by patient ID, StudyInstance UID, and ImageSeries UID, and compresses it into a zip file. When caIntegrator notifies you that the file is retrieved, the DICOM Retrieval page indicates whether the retrieved files are Study Instance UIDs or Image Series UIDs, shown in the following figure.
Click the Download DICOM link to download and save the file. caIntegrator unzips the file and displays the list of images in the file. To open the DICOM images, you must have a DICOM image viewer application installed on your computer. For more information about one such viewer, see http://www.e-dicom.com/viewers.php.
In the search results, not all of the subjects in the data subset may be mapped to image series IDs. If you select a mixture of subjects, some of which have image annotations as indicated by an image series ID and some of which do not have image annotations (no image series ID), when you click the Retrieve DICOM Images button, NBIA retrieves the images for the entire NBIA study instance UID that includes the image seriesIDs you checked.
If on the Search Results tab you select only subjects that have image annotations as indicated by an image series ID, when you click the Retrieve DICOM Images button, NBIA retrieves images for the NBIA image series that were matched in the search. If the results are a mixture, but you select one specific row with a valid image annotation, caIntegrator aggregates to the image series. If results are a mixture and you select multiple rows, caIntegrator aggregates to the NBIA study in which multiple image series you have selected in the search results are found.
If your query does not have image annotations and all check boxes are selected, results will go up to image series UID and gives all image series in it. Search results may ultimately depend on how the study was created. For example, if no image series display in query results, it means they were not mapped in the study. In that case, the results "move" up to Study Instance UIDs.
To best understand this, it is important to review the hierarchy of submissions in NBIA. For more information, see #Relationship of Subject to Study to Series to Images.
If you are searching a study that has image data and image annotation(s) for at least one image series, you would follow these steps:
When the image Study is in the checked boxes (regardless of image series being there or not), the system aggregates up to the Image Study level.
You can choose to download tabular search results as a CSV file. Click the Export .csv link at the bottom of the page. You may need to scroll the page to see it. The file contains the annotations, columns and data sort configurations you specified in the search query.
You will not see the Export option when gene expression data displays as query results. |