- Documentation Main Page
- Creation of the Cancer Gene Index
- Data, Metadata, and Annotations
- Cancer Gene Index Gene-Disease and Gene-Compound XML Documents
- caBIO APIs
- Cancer Gene Index Shared Parsed Data and Code
- caBIO Portlet Templated Searches
- caBIO Home Page
- caBIO iPhone Application
- caBIO Portlet Simple Searches
- Glossary
- Credits and Resources
Need Additional Help?
If you need additional support, please contact Application Support.
To Print the Guide
We recommend you print one wiki page of the guide at a time. To do this, click the printer icon at the top right of the page; then from the browser File menu, choose Print. Printing multiple pages at one time is more complex. For instructions, refer to How do I print multiple pages?.
Having Trouble Reading the Text?
Resizing the text for any web page is easy. For information on how to do this in your web browser, refer to this W3C tutorial .
Search for Biological Entities Disease Query Overview
The caBIO Home Page Search for Biological Entities is a flexible tool, and the workflow presented here is just one of the ways that it can be used. These steps illustrate how to query a specific type object to identify those with attributes that are associated with your disease search term.
Identifying a Disease Search Term
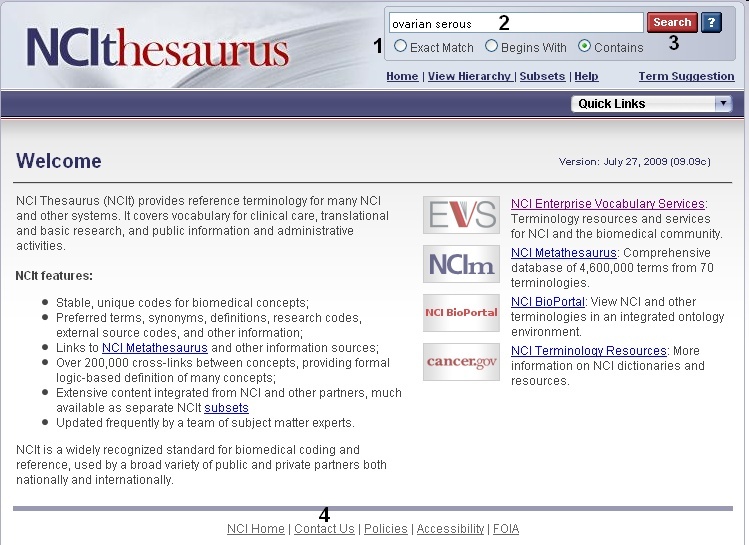
In order to search for genes that are associated with a given disease, you must first have either an NCI Thesaurus disease term or concept code. To find both of these values, navigate to the NCI Thesaurus web page. Select the Contains radio button (1). Enter your keyword/s (2), and click the Search button (3). If you cannot find the term for which you were looking, click on the Contact Us link at the bottom of the web page (4).
Tip
You may use the question mark wild card character after your search term (for example, "ovarian?") if you choose to instead use Exact Match or Begins With.
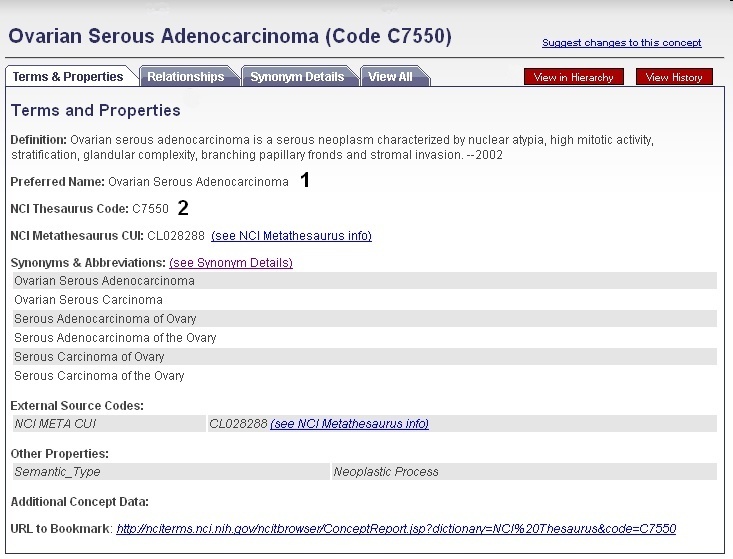
Select the desired result from the list of retrieved thesaurus terms in order to view the term's concept page. You may use either the Preferred Term (1) or the Thesaurus Code (2) with the Search for Biological Entities tool.
Tip
The NCI Thesaurus Code for a concept is identical to its EVS Identifier.
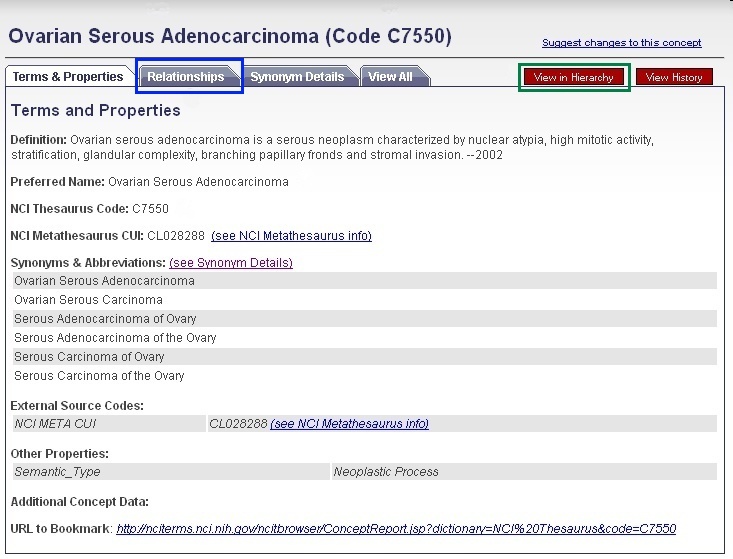
If you would like to uncover disease ontologies and also query the Search for Biological Entities functionality for parent and child concepts to your desired term, you may find the names and NCI Thesaurus Codes for the immediate parent or child concepts by clicking on the Relationships tab (blue box). Alternatively, you may view parent, sister, and child disease concepts by clicking View in Hierarchy (green box).
Using the Search for Biological Entities tool
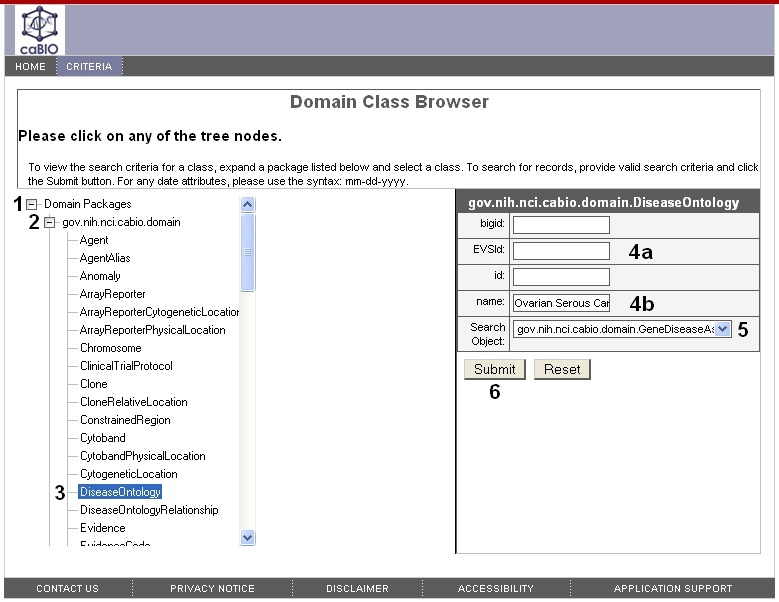
To search for genes that are associated with a particular disease, in the Domain Packages tree (1) expand the gov.nih.nci.cabio.domain package tree node (2) and select DiseaseOntology class (3). In the gov.nih.nci.cabio.domain.DiseaseOntology search form, provide the disease term NCI Thesaurus concept code in the EVSid field (4a) or the disease NCI Thesaurus term name in the name field (4b). Select the {{gov.nih.nci.cabio.domain.GeneDiseaseAssociation from the Search Object drop-down list (5), and click the Submit button (6).
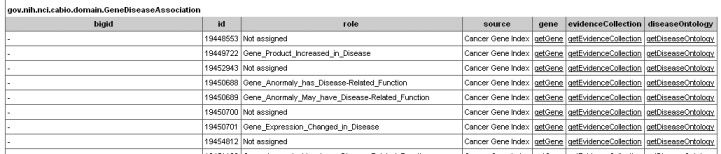
Retrieved results are listed as records in a single table (top panel), which includes each object's identifier, the Role Code and/or Role Detail associated with the gene-disease concept pair, notation that the data derive from the Cancer Gene Index, and three method links - getGene, getEvidenceCollection, and getDiseaseOntology.
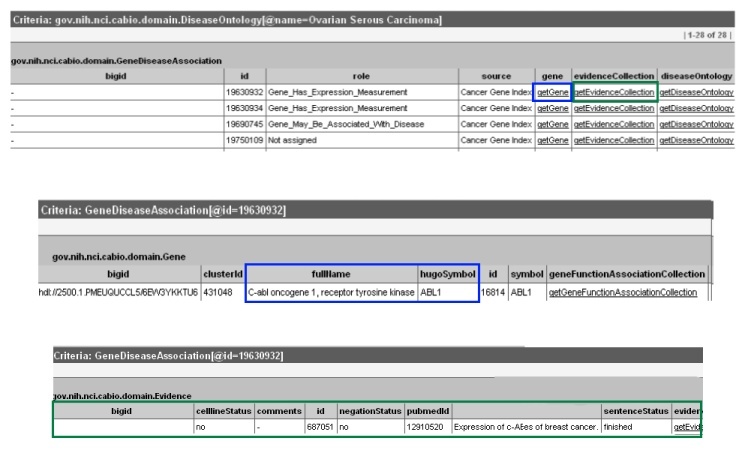
To view the gene-disease sentence and annotation information, select the getEvidenceCollection method (green box in top panel) to call up the associated Evidence type object (green box in middle panel). For more information on these data, refer to the Data, Metadata, and Annotations section.
Search Tip
If you do not want to spend time navigating through the caBIO object model for candidate gene-disease associations that were found to be false positives by expert human curators, you should first view the Evidence objects by clicking getEvidenceCollection (bottom panel). Scroll to the right to check that the Sentence Status is finished and Negation Status is no before clicking getGene on the results page.
Click on the getGene method link of any record (blue box in top panel) to access the related Gene object, which contains the full name and HUGO Gene Symbol in the fullName and hugoSymbol columns for the gene associated with the evidence and disease of interest (blue box in middle panel).
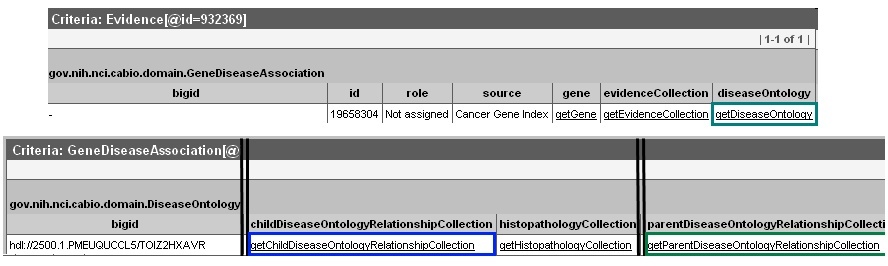
Click the getDiseaseOntology method link in any retrieved result should you wish to view parent and child concepts for your disease of interest and explore gene-disease associations with those disease concepts, as well (2). You can find parent disease concepts by scrolling to the right and selecting the getParentDiseaseOntologyRelationshipCollection link; child disease concepts can be accessed by clicking on the getChildDiseaseOntologyRelationshipCollection link.
Note
If the disease concept of interest has neither parent nor child concepts, you must search the NCI Thesaurus using your disease term or EVS Identifier listed in the Disease Ontology object.