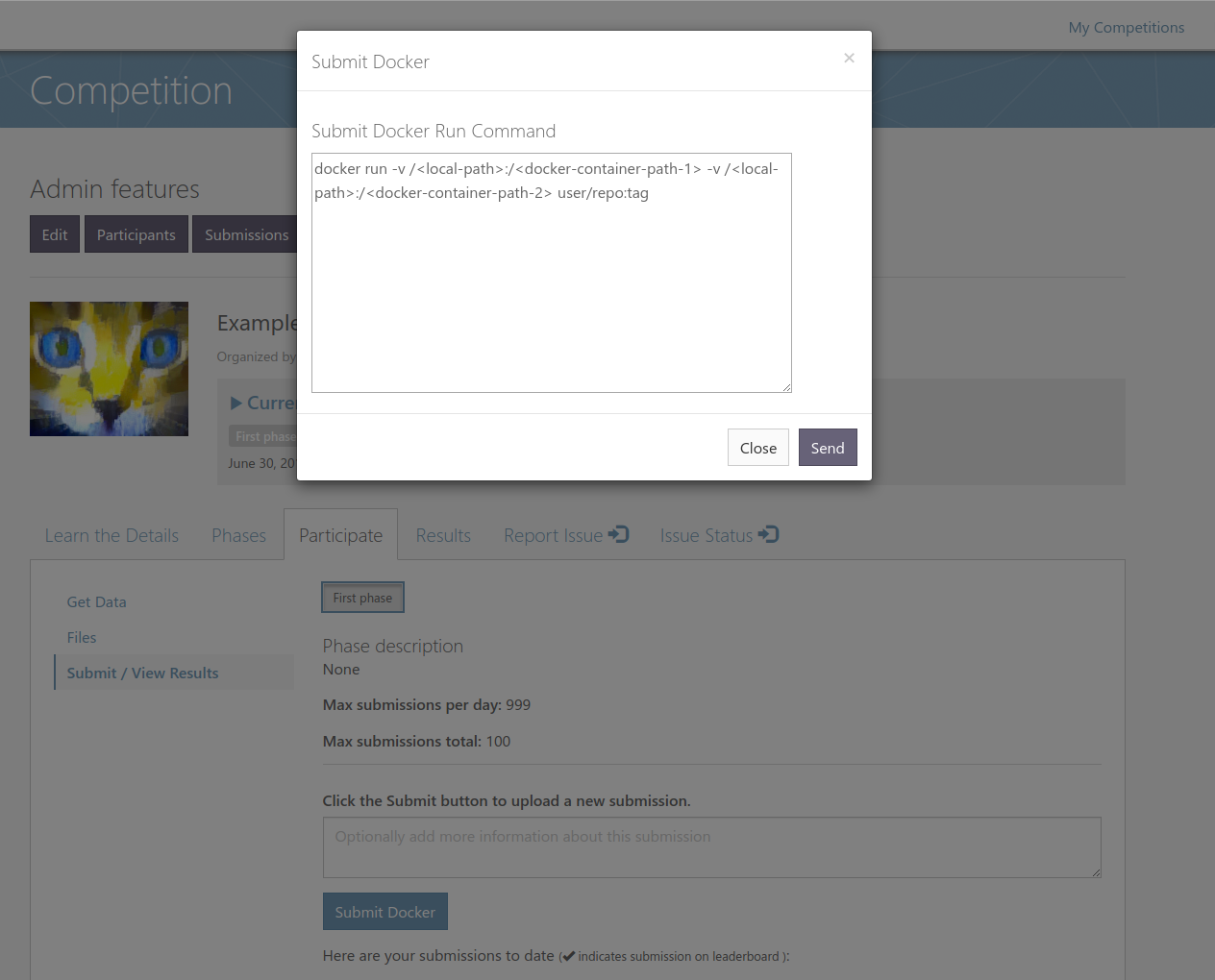
1) Re-implemented docker submissions for MedICI site (should work for upcoming challenges).
2) Got a tutorial for ePAD and learned how to make annotations and save them to the DB.
3) Also learned to download TCIA images and add them to ePAD.
| Planned for next month |
|---|
1) Fine tune code changes. 2) QA implementation by re-creating MICCAI 2019 challenge and submitting dockers to test implementation. |
| Comments |
|---|
Things that went well:
Things that could be improved:
|
| Meeting | Date |
|---|---|
| Bi-weekly Meeting #1 |
|
| Bi-weekly Meeting #2 |
|
Milestones
| Description | Date |
|---|---|
| Successfully got a docker run command to a worker VM and executed it. | 01/06/2019 |
| Learned enough about ePAD to setup ground truth annotation sets for professional annotators. |
Task
| Description | Resolution | Status | Creation Date | Close Date |
|---|---|---|---|---|
| Figure out how to run a docker through CodaLab | Complete | Done | 10/09/2019 | 01/06/2019 |
| Finish ePAD tutorials (they are online)..there is always more to learn | In Progress | In progress | 10/09/2019 | -- |
| Contact Ryan Birmingham for caMicroscope tutorial | In Progress | Coordinating with Ryan | 10/09/2019 | -- |
Risks
| Description | Mitigation | Rank | Status | Creation Date | Realization | Close Date |
|---|---|---|---|---|---|---|
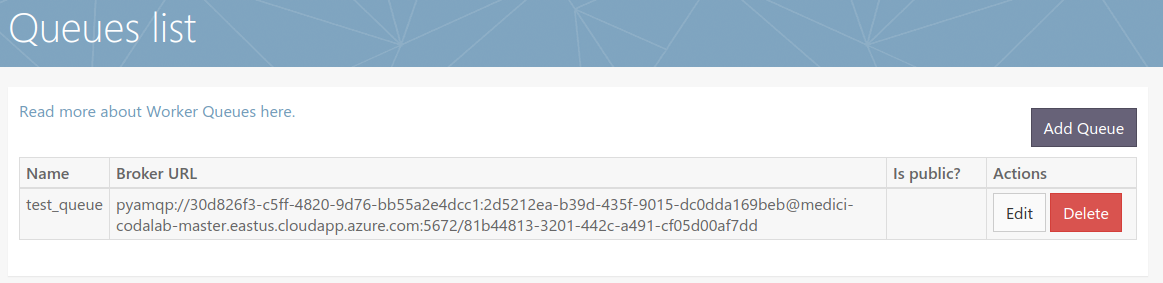
| The worker queue system in theory can handle multiple submissions by participants. Should that assumption not hold we could be in for a nasty surprise that this doesn't work as expected. | Test real AI containers that can take up to half a day to run on the system and submit multiple containers. | 1 | In Progress | 1/13/2020 | 1/13/2020 | -- |