|
Page History
...
Figure 5 Portal for Kaggle, a leading website for challenges for data scientists
Topcoder (https://www.topcoder.com) is a similar popular website for software developers, graphic designers and data scientists. In this case, participants typically share their code or designs. They use the Appirio proprietary crowdsourcing development platform, built on Amazon Web Services, Cloud Foundry, Heroku, HTML5, Ruby and Java. A recent computational biology challenge run on Topcoder demonstrated that this crowdsourcing approach produced algorithmic solutions that greatly outperform commonly used algorithms such as BLAST for sequence annotation {Lakhani, 2013 #3789}. This competition was run with a $6000 prize and drew 733 participants (17% of whom submitted code) and the prize-winning algorithms were made available with an open source license.
Challenge Post (http://challengepost.com/ ) has been used to organize hackathons, online challenges and other software collaborative activities. In person hackathons are free while the online challenges cost $1500/month (plus other optional chargesWhere does the section for commercial challenge management systems end?Could the suitability of each platform for imaging challenges be discussed?charges).
Open Source
Synapse is both an open source platform and a hosted solution for challenges and collaborative activities created by Sage bionetworks. It has been used for a number of challenges including the DREAM challenge. Synapse allows the sharing of code as well as data. However, the code typically is in R, Python and similar languages. Synapse also has a nice programmatic interface and methods to upload/download data, submit results, create annotations and provenance through R, Python, command line and Java. These options can be configured for the different challenges. Content in Synapse is referenced by unique Synapse IDs. The three basic types of Synapse objects include projects, folders and files. These can be accessed through the web interface or through programmatic APIs. Experience and support for running image analysis code within Synapse is limited.
...
Figure 7 Example Challenge hosted in Synapse
COMIC framework (http://grand-challenge.org/) is an open-source platform that facilitates the creation of challenges and has been used to host a number of medical imaging challenges. The Consortium for Open Medical Image Computing (COMIC) platform, built using Python/Django was created and is maintained by a consortium of five European medical image analysis groups including Radboud University, Erasmus, and UCL. They also offer a hosted site, with the hardware located at Fraunhofer MEVIS in Bremen, Germany. The current framework allows participants to create a website, add pages including wikis, create participant registrations, methods for organizers to upload data and participants to download data (for instance through Dropbox). However, the platform including ways to visualize medical data and results is still under development as are options to share algorithms and perform challenges in the cloud.
The main steps to create a new challenge are http://grand-challenge.org/support/ :
- Create a project
- Add pages
- Making uploaded files available for download
- Allowing others to register for your project
- Make your project appear in the projects overview
- Allow file uploads
- Including content from files on a page
- Allow others to edit the project
- Changing colors and other styling
- Project data folder
- Page permissions
However, at this time, there is limited support for automatic evaluation of submitted results, results presentation, native support for medical images although many of these features are planned.
The HubZero (https://hubzero.org/) is an open source platform developed for scientific collaboration. It has been used heavily in a number of communities including nanoscience, earthquake engineering, molecular diagnostics and others. A version focused on cancer informatics can is hosted at nciphub.org. nciphub shares a lot of features with the Synapse platform. It allows user management and role-based access. Users can create groups that share common interest and collaborate within these groups. Files can be shared within projects. Other features include wiki, calendars, creating and sharing resources such as presentations, multimedia and even tools. Most common tools found on the various hubs are those based on simulations. Although nciphub has limited native support of medical imaging, libraries to handle medical images can be configured to work in the hub. Members of the Quantitative Imaging Network (QIN) are exploring the use of nciphub for challenges, especially for the communication and data sharing.
CodaLab (http://codalab.org/) is an open-source project that originated at Microsoft Research that was expressly created for hosting challenges and supporting reproducible research. The OuterCurve Foundation currently maintains it. Challenge organizers can easily set up challenges by creating a competition bundle that consists of data as well as evaluate tools. As part of the configuration files, the number of phases and duration (e.g. training, leaderboard, test) can be set up by the organizer. The evaluation program can be written in any language. Participants can upload results and get immediate feedback. The currently available version of CodaLab comes with scoring algorithms for image segmentation evaluation Organizers can extend the presentation of results to allow drilling down into the results with tables and charts. CodaLab currently uses the Azure platform although, in theory, it should be possible to deploy on other servers without a great deal of effort. CodaLab is also developing support for worksheets. These are resources to support reproducible research and for collaboration. Using these, researchers have compared a number of open source NLP tools on different public datasetshttp://research.microsoft. com/apps/video/default.aspx?id=225950 . As this technology continues to be developed, researchers will be able to quickly compare the performance of different algorithms on a range of datasets in the "cloud" by leveraging Azure technology.
...
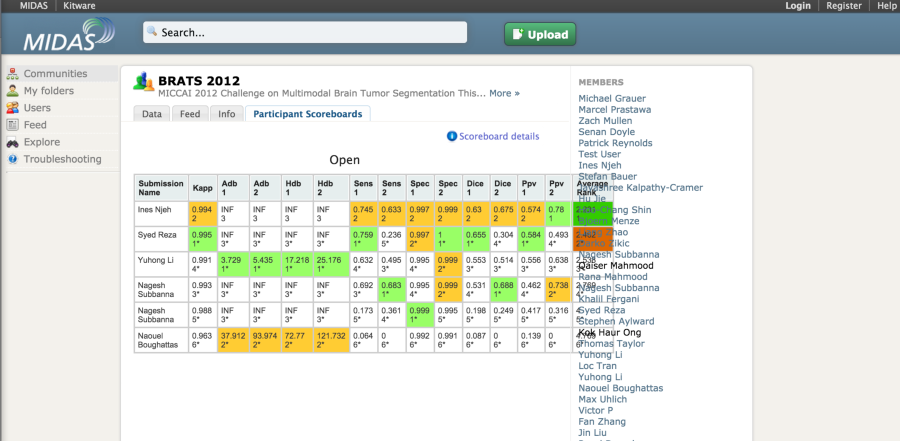
The MIDAS platform has been used to host a couple of imaging challenges. A special module is available to host challengeshttps://github. com/midasplatform/challenge . The developers of the platform also made available the COVALIC evaluation tool for segmentation challenges with the following metrics: Average distance of boundary surfaces, 95th percentile Hausdorff distance of boundary surfaces, Dice overlap, Cohen's kappa, Sensitivity, Specificity, Positive Predictive Value.
Once the participants have uploaded their submissions, the leaderboard updates the scores automatically.
A new version of the platform appears to be in development. http://challenge.kitware. com/#challenges/learn This system (COVALIC) is built on the TangeloHub platform--an open source data and analytics platform made up of three major components: Tangelo, Girder, and Romanesco.
...