|
Page History
| Scrollbar | ||
|---|---|---|
|
| Page info | ||||
|---|---|---|---|---|
|
This chapter describes how to use caIntegrator tools to analyze data in subject annotation or genomic studies that have been deployed in caIntegrator.
...
| Table of Contents | ||
|---|---|---|
|
Data Analysis Overview
Once a study has been deployed, you can analyze the data using caIntegrator analysis tools.
...
- K-M Plot: This tool analyzes subject annotation data, generating a Kaplan-Meier (K-M) plot based on survival data sets. See #Creating Kaplan-Meier Plots.
- Gene Expression Plot: This tool analyzes annotation, subject annotation or genomic data based on gene expression values. See #Creating Gene Expression Plots.
- GenePattern: This feature provides an express link to GenePattern where you can perform analyses on selected caIntegrator studies, or it enables you to perform several GenePattern analyses on the grid. See #Analyzing Data with GenePattern.
After defining or running the analysis on selected data sets, analysis results display on the same page, allowing you to review the analysis method parameters you defined.
Creating Kaplan-Meier Plots
This help topic opens from any of the three K-M plot tabs. For specific details about working with these tabs, see the following topics:
The Kaplan-Meier method analyzes comparative groups of subjects or samples. In caIntegrator, the K-M method can compare survival statistics among comparative groups. You can configure the survival data in the application. For example, you might identify a group of patients with smoking history and compare survival rates with a group of non-smoking patients, or compare the survival data for two groups of patients with a specific disease type, based on Karnofsky scores. You could compare groups of subjects with varying gene expression levels. You can also identify data sets using the query feature in the application, saving the queries, then configuring the K-M to compare groups identified by the queries.
...
| Info | ||
|---|---|---|
| ||
To perform a K-M plot analysis, survival data must have been identified for the study you want to analyze or an Annotation Field Descriptor such as DAYSTODEATH has been set to Data Type 'numeric'. For more information, see Defining Survival Values. |
K-M Plot for Annotations
The groups identified for this K-M plot generation are based on annotations.
...
- The number of subjects for each group is embedded in the legend below the plot.
- caIntegrator generates a P-value for the selected groups; it displays at the bottom of the page. A low P-value generally has more significance than a high P-value.
- For information regarding the P-value calculation, see #Creating Kaplan-Meier Plots.
K-M Plot for Gene Expression
caIntegrator allows you to compare expression levels for one given gene in different representative groups. The relative expression level is referred to as "fold change". Fold change is the ratio of the measured gene expression value in an experimental sample as determined by a reporter to a reference value calculated for that reporter against all control samples. The reference value is calculated by taking the mean of the log2 of the expression values for all control samples for the reporter in question. The log2 mean value, n, is then converted back to a comparable expression signal by returning 2 to the exponent n.
...
- The gene symbol for each group represented in the data appears with its color correlation to the plot embedded in the legend below the plot. Three lines on this plot represent each gene symbol entered for the plot. Each line of the three represents a subgroup of people carrying the gene--one line for overexpressed values, one line for under expressed values and one line for intermediate values which represents gene values that are not up-regulated nor down-regulated.
- In queries that include a fold change criterion and that are configured to return genomic data, raw expression values are replaced with calculated fold change values.
- A P-value is also generated for the selected groups; it displays at the bottom of the page. A low P-value generally has more significance than a high P-value.
- For information regarding the P-value calculation, see #Creating Kaplan-Meier Plots.
K-M Plot for Queries and Saved Lists
You can identify data sets using the query feature in the application. You can manipulate the queries to find the groups you want to compare, save the queries, then configure the K-M to compare the query groups. This is one method of limiting the data considered in the K-M plot calculation.
...
- The number of subjects for each group is embedded in the legend below the plot.
- A P-value is also generated for the selected groups; it displays at the bottom of the page. A low P-value generally has more significance than a high P-value.
- For information regarding the P-value calculation, see #Creating Kaplan-Meier Plots.
Creating Gene Expression Plots
Gene expression plots compare signal values from reporters or genes. This statistical tool allows you to compare values for multiple genes at a time; it does not limit your comparison to only two sets of data. It also allows you to compare expression levels for selected genes against expression levels for a set of control samples designated at the time of study definition.
caIntegrator provides three ways to generate meaningful gene expression plots, indicated by tabs on the page. The tabs are independent of each other and allow you to select the genes, reporters and sample groups to be analyzed on the plot.
- #Gene Expression Value Plot for Annotation – You can locate genes in the caBIO directories or caIntegrator Gene Lists. You can learn more about the genes in the CGAP directory. You can define criteria for the plot using subject annotation and image annotations.
- #Gene Expression Value Plot for Genomic Queries – You can select data based on saved genomic queries.
- #Gene Expression Value Plot for Annotation and Saved List Queries – You can select data based on saved subject annotation queries. You can locate genes in the caBIO directories or caIntegrator Gene Lists.
See also #Understanding a Gene Expression Plot.
Gene Expression Value Plot for Annotation
To generate a gene expression plot, follow these steps:
...
- You can recalculate the data display by changing the Plot Type above the graph.
- You can modify the plot parameters and click the Reset button to recalculate the plot.
See also See #Understanding a Gene Expression Plot.
Gene Expression Value Plot for Genomic Queries
Data to be analyzed on this tab must have been saved as a genomic query. For more information, see Saving a Query.
...
- You can recalculate the data display by changing the Plot Type above the graph.
- You can modify the plot parameters and click the Reset button to recalculate the plot.
See also #Understanding a Gene Expression Plot.
Gene Expression Value Plot for Annotation and Saved List Queries
Data to be analyzed on this tab must have been saved as a subject annotation query, but it must have genomic data identified in the query. For more information, see Adding/Editing Genomic Data. For the genomic data, you must identify genes whose expression values are used to calculate the plot.
...
- You can recalculate the data display by changing the Plot Type above the graph.
- You can modify the plot parameters and click the Reset button to recalculate the plot.
See also See #Understanding a Gene Expression Plot.
Understanding a Gene Expression Plot
Above the plot, you can select various plot types. When you do so, the plot is recalculated. Although all of the plots in this section appear similar, note the differences in calculation results and legends between the Y axis on each of the plots.
...
| Include Page | ||||
|---|---|---|---|---|
|
Analyzing Data with GenePattern
GenePattern is an application developed at the Broad Institute that enables researchers to access various methods to analyze genomic data. caIntegrator provides an express link to GenePattern where you can analyze data in any caIntegrator study.
...
The GenePattern feature in caIntegrator currently supports three analyses on the grid: #Comparative Marker Selection (CMS) Analysis, #Principal Component Analysis (PCA) and #GISTIC-Supported Analysis.
| Tip | ||
|---|---|---|
| ||
If you are using the web interface to access GenePattern (option #1 listed above), then you can run other GenePattern tools in addition to CMS, PCA and GISTIC. |
- Select the study whose data you want to analyze in the upper right portion of the caIntegrator page.
- Click GenePattern Analysis in the left sidebar of caIntegrator. This opens the GenePattern Analysis Status page, shown in the following figure.
- Select from the drop-down list the type of GenePattern analysis you want to run on the data.
- GenePattern Modules--This option launches a session within GenePattern from which you can launch analyses. See #Gene Pattern Modules.
- Comparative Marker Selection (Grid Service)--This option enables you to run this GenePattern analysis on the grid. See #Comparative Marker Selection (CMS) Analysis.
- Principal Component Analysis (Grid Service). This option enables you to run this GenePattern analysis on the grid. See #Principal Component Analysis (PCA).
- GISTIC (Grid Service). This option enables you to run this GenePattern analysis on the grid. See #GISTIC-Supported Analysis|.
- GenePattern Modules--This option launches a session within GenePattern from which you can launch analyses. See #Gene Pattern Modules.
- Click the New Analysis Job button to open a corresponding page where you can configure the analysis parameters.
GenePattern Modules
| Info | ||
|---|---|---|
| ||
To launch the analyses described in this section, you must have a registered GenePattern account. For more information, see http://genepattern.broadinstitute.org/gp/pages/login.jsf |
- To configure the link for accessing GenePattern from caIntegrator, open the appropriate page as described in #Analyzing Data with GenePattern.
- Select the study whose data you want to analyze in the upper right portion of the caIntegrator page.
- Click GenePattern Analysis in the left sidebar of caIntegrator. This opens the GenePattern Analysis Status page.
- Make sure GenePattern Modules is selected in the drop down list. Click New Analysis Job.
- In the GenePattern Analysis dialog box, shown in the following figure, specify connection information and click Connect. Fields are described in the table following the figure.
Field
Description
Server URL
Enter any GenePattern publicly available URL, such as http://genepattern.broadinstitute.org/gp/services/Analysis.
GenePattern Username
Enter your GenePattern user name.
GenePattern Password
Enter your GenePattern password.
- After logging in with the GenePattern profile, the dialog box, shown in the following figure, expands to include fields for defining the GenePattern analysis.
- Enter information for the following fields. Fields with a red asterisk are required.
Field
Description
Job Name*
Enter a unique name for the analysis
Analysis Method
Select any method from the drop down list. Click Analysis Method Documentation for descriptions of the different analysis methods.
Data
All genomic data is selected by default. Select from the list any list that has been created for this study.
cls
Select any annotation field.
The CLS file format defines phenotype (class or template) labels and associates each sample in the expression data with a label. It uses spaces or tabs to separate the fields. The CLS file format differs somewhat depending on whether you are defining categorical or continuous phenotypes:
Categorical labels define discrete phenotypes; for example, normal vs tumor).
Continuous phenotypes are used for time series experiments or to define the profile of a gene of interest (gene neighbors).Info title Note Most GenePattern modules are intended for use with categorical phenotypes. Therefore, unless the module documentation explicitly states otherwise, a CLS file should define categorical labels.
prediction.results.file
Enter the name of this file which is part of the output from a GenePattern module.
- Click Perform Analysis. Based on the analysis method you select, you may be asked to add more information for the analysis. For more information, refer to the GenePattern Help site: http://genepattern.broadinstitute.org/gp/getTaskDocCatalog.jsp.90
...
You can run GenePattern analyses for Comparative Marker Selection, Principal Component Analysis and GISTIC-based analysis on the grid if you choose.
Comparative Marker Selection (CMS) Analysis
The Comparative Marker Selection (CMS) module implements several methods to look for expression values that correlate with the differences between classes of samples. Given two classes of samples, CMS finds expression values that correlate with the difference between those two classes. If there are more than two classes, CMS can perform one-vs-all or all-pairs comparisons, depending on which option is chosen.
...
- Select the study whose data you want to analyze in the upper right portion of the caIntegrator page. You must select a study saved as a subject annotation study, but which has genomic data.
- Click GenePattern Analysis in the left sidebar of caIntegrator. This opens the GenePattern Analysis Status page.
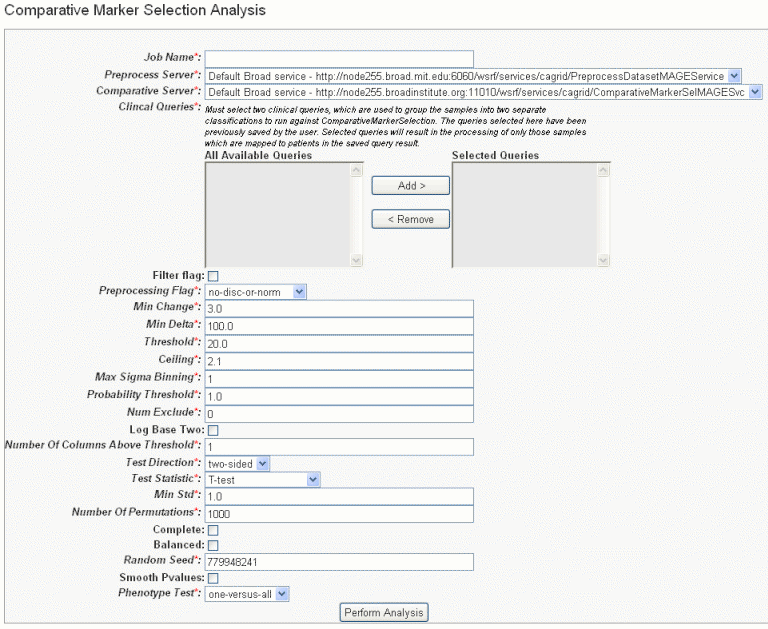
- In the GenePattern Analysis Status page, select Comparative Marker Selection (Grid Service) from the drop down list and click New Analysis Job. This opens the Comparative Marker Selection Analysis page, shown in the following figure.
- Select or define CMS analysis parameters, described in the following table. An asterisk indicates required fields. The default settings are valid; they should provide valid results.
CMS Parameter
Description
Job Name*
Assign a unique name to the analysis you are configuring.
Preprocess Server*
A server which hosts the grid-enabled data GenePattern PreProcess Dataset module. Select one from the list and caIntegrator will use the selected server for this portion of the processing.
Comparative Server*
A server which hosts the grid-enabled data GenePattern Comparative Marker Selection module. Select one from the list and caIntegrator will use the selected server for this portion of the processing.
Annotation Queries and Lists*
All subject annotation queries and gene lists with appropriate data for the analysis are listed. Select and move two or more queries from the All Available Queries panel to the Selected Queries panel using the Add > and Remove < buttons.
<ac:structured-macro ac:name="unmigrated-wiki-markup" ac:schema-version="1" ac:macro-id="715195b64cc34647-6fb0b6bd-42954592-8a9d82a2-d3ca59b0b600e9ab31430e65"><ac:plain-text-body><![CDATA[Note: The [SL] and [Q] prefixes to list names indicate "Subject Lists" or "Saved Queries". A "G" in the prefix indicates the list is Global. For more information, see [Creating a Gene or Subject Listhttps://wiki.nci.nih.gov/x/FoDnAg#4-ViewingQueryResults-CreatingaGeneorSubjectList].
]]></ac:plain-text-body></ac:structured-macro>
Filter Flag
Variation filter and thresholding flag
Preprocessing Flag*
Discretization and normalization flag
Min Change*
Minimum fold change for filter
Min Delta*
Minimum delta for filter
Threshold*
Value for threshold
Ceiling*
Value for ceiling
Max Sigma Binning*
Maximum sigma for binning
Probability Threshold*
Value for uniform probability threshold filter
Num Exclude*
Number of experiments to exclude (max & min) before applying variation filter
Log Base Two
Whether to take the log base two after thresholding; default setting is "Yes".
Number of Columns Above Threshold*
Remove row if n columns are not >= than the given threshold
In other words, the module can remove rows in which the given number of columns does not contain a value greater or equal to a user defined threshold.Test Direction*
The test to perform (up-regulated for class0; up-regulated for class1, two sided). By default, Comparative Marker Selection performs the two-sided test.
Test Statistic*
Select the statistic to use.
Min Std*
The minimum standard deviation if test statistic includes the min std option. Used only if test statistic includes the min std option.
Number of Permutations*
The number of permutations to perform. (Use 0 to calculate asymptotic P-values.) The number of permutations you specify depends on the number of hypotheses being tested and the significance level that you want to achieve (3). The greater the number of permutations, the more accurate the P-value.
Complete – Perform all possible permutations. By default, complete is set to No and Number of Permutations determines the number of permutations performed. If you have a small number of samples, you might want to perform all possible permutations.
Balanced – Perform balanced permutationsRandom Seed*
The seed for the random number generator.
Smooth P-values
Whether to smooth P-values by using the Laplace's Rule of Succession. By default, Smooth P-values is set to Yes, which means P-values are always less than 1.0 and greater than 0.0.
Phenotype Test*
Tests to perform when class membership has more than 2 classes: one versus-all, all pairs.
Note: The P-values obtained from the one-versus-all comparison are not fully corrected for multiple hypothesis testing. - When you have completed the form, click Perform Analysis.
caIntegrator takes you to the JobStatus/Launch page where you will see the job and its status in the Status column of the list, shown in the following figure.
- When the job is complete, the system displays a completion date on the GenePattern Analysis status page. Click the Download link. This downloads zipped result files to your local work station. The number of files and their file type will vary according to the processing. The results format is compatible with GenePattern visualizers and can be uploaded within GenePattern.
Principal Component Analysis (PCA)
Principal Component Analysis is typically used to transform a collection of correlated variables into a smaller number of uncorrelated variables, or components. Those components are typically sorted so that the first one captures most of the underlying variability and each succeeding component captures as much of the remaining variability as possible.
...
- Select the study whose data you want to analyze in the upper right portion of the caIntegrator page. You must select a study with gene expression data.
- Click GenePattern Analysis in the left sidebar of caIntegrator. This opens the GenePattern Analysis Status page.
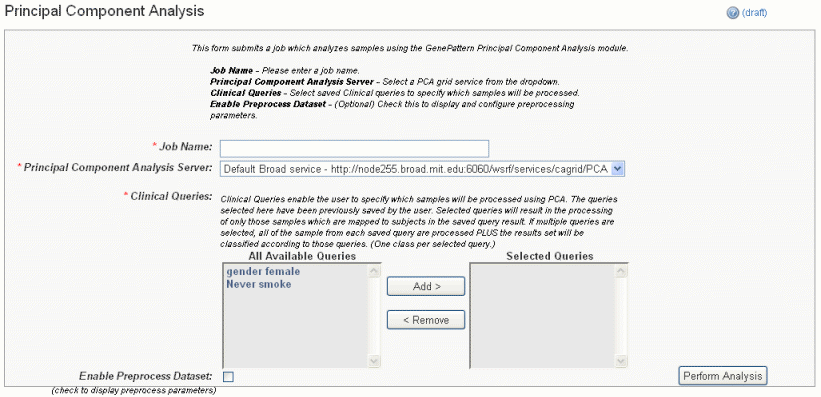
- Select Principal Component Analysis (Grid Service) from the drop down list and click New Analysis Job. This opens the Principal Component Analysis page, shown in the following figure.
- Select or define PCA analysis parameters, described in the following table. An asterisk indicates required fields. You must enter a job name and select an annotation query, but you can accept the default settings for other options.
PCA Parameters
Description
Job Name*
Assign a unique name to the analysis you are configuring.
Principal Component Analysis Server*
A server which hosts the grid-enabled data GenePattern Principal Component Analysis module. Select one from the list and caIntegrator will use the selected server for this portion of the processing.
Annotation Queries*
All annotation queries display in this list. Select one or more of these queries to define which samples are analyzed using PCA. If you select more than one query, then the union of the samples returned by the multiple queries is analyzed.
Cluster By*
Selecting rows looks for principal components across all expression values, and selecting columns looks for principal components across all samples.
- If you want to preprocess the data set, click Enable the Preprocess Dataset. This opens an additional set of parameters, shown in the following figure and described in the following table. The preprocessing is executed prior to running the PCA.
PCA Preprocessing Parameters
Description
Preprocess Server*
A server which hosts the grid-enabled data GenePattern PreProcess Dataset module. Select one from the list and caIntegrator will use the selected server for this portion of the processing.
Filter Flag
Variation filter and thresholding flag
Preprocessing Flag
Discretization and normalization flag
Min Change
Minimum fold change for filter
Min Delta
Minimum delta for filter
Threshold
Value for threshold
Ceiling
Value for ceiling
Max Sigma Binning
Maximum sigma for binning
Probability Threshold
Value for uniform probability threshold filter
Num Exclude
Number of experiments to exclude (max & min) before applying variation filter
Log Base Two
Whether to take the log base two after thresholding
Number of Columns Above Threshold
Remove row if n columns no >= than the given threshold
- When you have completed the form, click Perform Analysis.
- When the job is complete, the system displays a completion date on the GenePattern Analysis status page. Click the Download link. This downloads zipped result files to your local work station. The number of files and their file type will vary according to the processing. The results format is compatible with GenePattern visualizers and can be uploaded within GenePattern.
GISTIC-Supported Analysis
| Info | ||
|---|---|---|
| ||
The GISTIC test option displays only if the study contains copy number or SNP data. For more information, see Configuring Copy Number Data. |
...
- Select the study whose data you want to analyze in the upper right portion of the caIntegrator page. You must select a study with copy number (either Affymetrix SNP or Agilent Copy Number) data.
- Click GenePattern Analysis in the left sidebar of caIntegrator. This opens the GenePattern Analysis Status page.
- In the GenePattern Analysis Status page, select GISTIC (Grid Service) from the drop down list and click New Analysis Job. This opens the GISTIC Analysis page, shown in the following figure.
- Select or define GISTIC analysis parameters, described in the following table. You must indicate a Job Name, but you can accept the other default settings, which are valid and should produce valid results. Asterisks identify required fields.
GISTIC Parameters
Description
Job Name*
Assign a unique name to the analysis you are configuring.
GISTIC Service Type*
Select whether to use the GISTIC web service or grid service and provide or select the service address. If the web service is selected, authentication information is also required
GenePattern User Name/Password
Include these to log into GenePattern for the analysis.
Annotation Queries and Lists
All annotation queries display in this list as well as an option to select all non-control samples. Select an annotation query if you wish to run GISTIC on a subset of the data and select all non-control samples if wish to include all samples.
Select Platform
This option appears only if more than one copy number platform exists in the study. Select the appropriate platform from the drop-down list ().
Exclude Sample Control Set*
From the drop-down list, select the name of the control set you want to exclude from the analysis. Click None if that is applicable.
Amplifications Threshold*
Threshold for copy number amplifications. Regions with a log2 ratio above this value are considered amplified. Default = 0.1.
Deletions Threshold*
Threshold for copy number deletions. Regions with a log2 ratio below the negative of this value are considered deletions. Default = 0.1.
Join Segment Size*
Smallest number of markers to allow in segments from the segmented data. Segments that contain fewer than this number of markers are joined to the neighboring segment that is closest in copy number. Default = 4.
<ac:structured-macro ac:name="unmigrated-wiki-markup" ac:schema-version="1" ac:macro-id="c1402912d2b3d27d-5867dcf5-4ac2489e-9302b8cc-450a09a00d78b74b975db1e1"><ac:plain-text-body><![CDATA[
QV Thresh[hold]*
Threshold for q-values. Regions with q-values below this number are considered significant. Default = 0.25.
]]></ac:plain-text-body></ac:structured-macro>
Remove X*
Flag indicating whether to remove data from the X-chromosome before analysis. Allowed values = {1,0}. Default = 1(yes).
cnv File
This selection is optional.
Browse for the file. There are two options for the CNV file.
Option #1 enables you to identify CNVs by marker name. Permissible file format is described as follows:
A two column, tab-delimited file with an optional header row. The marker names given in this file must match the marker names given in the markers_file. The CNV identifiers are for user use and can be arbitrary. The column headers are:
Marker Name
CNV Identifier
Option #2 enables you to identify CNVs by genomic location. Permissible file format is described as follows:
A 6 column, tab-delimited file with an optional header row. The 'CNV Identifier', 'Narrow Region Start' and 'Narrow Region End' are for user use and can be arbitrary. The column headers are:
CNV Identifier
Chromosome
Narrow Region Start
Narrow Region End
Wide Region Start
Wide Region End - When you have completed the form, click Perform Analysis.
- When the job is complete, the system displays a completion date on the GenePattern Analysis status page. Click the Download link. This downloads zipped result files to your local work station. The number of files and their file type will vary according to the processing. The results format is compatible with GenePattern visualizers and can be uploaded within GenePattern.
Additionally, upon completion of a successful GISTIC anaylsis, caIntegrator automatically displays the two gene lists that it generates in the Gene List Picker so that you can use them in a caIntegrator query or plot calculation. The lists are visible only to your userID. For more information, see #Choosing Genes. The genes will also display in Saved Copy Number Analyses in the left sidebar. See #Editing a GISTIC Analysis.
| Warning | ||
|---|---|---|
| ||
If samples from a copy number source are deleted, the GISTIC job in which they are appear is also deleted. |
Editing a GISTIC Analysis
To view a GISTIC analysis page in caIntegrator where you can review or edit analysis parameters and results, under Study Data in the left sidebar, click Saved Copy Number Analysis. Select the analysis you want to open. The system displays analysis parameters and gene lists that that were retrieved from the analysis, as shown in the following figure.
...