|
Page History
Infrastructure for Algorithm Comparisons, Benchmarks, and Challenges in Medical Imaging
AuthorAuthors: Jayashree Kalpathy-Cramer and Karl Helmer
| Panel | ||
|---|---|---|
| ||
|
...
Challenges are being increasingly viewed as a mechanism to foster advances in a number of domains, including healthcare and medicine. The US United States Federal governmentGovernment, as part of the open-government initiative, has underscored the role of challenges as a way to "promote innovation through collaboration and (to) harness the ingenuity of the American Public." Large quantities of publicly available data and cultural changes in the openness of science have now made it possible to use these challenges and crowdsourcing efforts to propel the field forward.
Sites such as Kaggle
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
In the biomedical domain, challenges have been used effectively in bioinformatics as seen by recent crowd-sourced efforts such as Critical Assessment of Protein Structure Prediction (CASP), the CLARITY Challenge for standardizing clinical genome sequencing analysis and reporting and the cancer Genome atlas Pan-cancer analysis Working Group, DREAM Challenges (
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
Some of the key advantages of challenges over conventional methods include 1) scientific rigor (sequestering the test data), 2) comparing methods on the same datasets with the same, agreed-upon metrics, 3) allowing computer scientists without access to medical data to test their methods on large clinical datasets, 4) making available resources resources available, such as source code, and 5) bringing together diverse communities (that may traditionally not work together) of imaging and computer scientists, machine learning algorithm developerdevelopers, software developers, clinicians, and biologists.
However, despite this potential, there are a number of challenges. Medical data is usually governed by privacy and security policies such as HIPPA that make it difficult to share patient data. Patient health records can be very difficult to completely deidentifyde-identify. Medical imaging data, especially brain MRIs, can be particularly challenging as once one could easily reconstruct a recognizable 3D model of the subject.
...
The medical imaging community has conducted a host of challenges at conferences such as MICCAI and SPIE. However, these have typically have been modest in scope (both in terms of data size and number of participants). Medical imaging data poses additional challenges to both participants and organizers. For organizers, ensure ensuring that the data are free of PHI is both critical and non-trivial. Medical data is typically acquired in DICOM format. However, ensuring that a DICOM file is free of PHI requires domain knowledge and specialized software tools. Multimodal imaging data can be extremely large. Imaging formats for pathology images can be proprietary and interoperability between formats can require additional software development efforts. Encouraging non-imaging researchers (e.g. machine-learning scientists) to participate in imaging challenges can be difficult due to the domain knowledge required to convert medical imaging into a set of feature vectors. For participants, access to large compute clusters with computing power, storage space, and bandwidth can prove difficult. Medical imaging data is challenging for non-imaging researchers.
However, it is imperative that the imaging community develops the tools and infrastructure necessary to host these challenges and potentially enlarge the pool of methods by making it more feasible for non-imaging researchers to participate. Resources such as the Cancer Imaging Archive (TCIA) have greatly reduced the burden for sharing medical imaging data within the cancer community and making these data available for use in challenges. Although a number of challenge platforms exist currently, we are not aware of any systems that meet all the requirements necessary to currently host medical imaging challengechallenges.
In this article, we review a few historical imaging challenges. We then list the requirements we believe to be necessary (and nice to have) to support large-scale multimodal imaging challenges. We then review existing systems and develop a matrix of features and tools. Finally, we make some recommendations for developing Medical Imaging Challenge Infrastructure (MedICI), a system to support medical imaging challenges.
...
Challenges have been popular in a number of scientific communities since the 1990s. In the text retrieval community, the Text REtrieval Conference (TREC), co-sponsored by NIST, is an early example of evaluation campaigns where participants work on a common task using data provided by the organizers and evaluated with a common set of metrics. ChaLearn
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
We begin with a brief review of a few medical imaging challenges held in the last decade and review their organization and infrastructure requirements. Medical imaging challenges are now a routine aspect of the highly regarded MICCAI annual meeting. Challenges at MICCAI began in 2007 with a liver segmentation and caudate segmentation challenges.
Grand Challenges in Biomedical Image Analysis maintains a fairly updated list of the
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
- A task is defined (the output). In our context, this could be segmentation of a lesion or organ, classification of an imaging study as being benign or malignant, prediction of survival, classification of a patients as being a responder or non-responder, pixel/voxel level classification of tissue or tumor grading.
- A set of images are provided (the input). These images are chosen to be of a sufficient size and diversity to reflect the challenges of the clinical problem. Data is typically spilt split up into training and test datasets. The "truth" is made available to the participants for the training data but not the test data. This reduces the risk of overfitting the data and ensures the integrity of the results.
- An evaluation procedure is clearly defined; given the output of an algorithm on a the test images, one or more metrics are computed that measure the performance, usually a reference output is used in this process, but it could also be a visual evaluation of the results by human experts)
- Participants apply their algorithm to all data in the public test dataset provided. They can estimate their performance on the training test.
- Some challenges have an optional leaderboard phase where a subset of the test images is made available to the participants ahead of the final test. Participants can submit their results to the challenge system and have them evaluated or ranked but these are not considered the final standing.
- The reference standard or "ground truth" is defined using methodology clearly described to the participants but is not made publicly available in order to ensure that algorithm results are submitted to the organizers for publication rather than retained privately.
- Final valuation is carried out by the challenge organizers on the test set where the ground truth is sequestered from the participants.
...
There were 3 sub-challenges within the radiology challenge. The primary goal of the radiology challenge was to perform segmentation from multimodal MRI of brain tumors. T1 (pre- and post constrast-contrast), T2 and FLAIR MRI images were preprocessed (registered and resampled to 1mm isotropic) by the organizers and made available. Ground truth in the form of label maps (4 color –enhancing, necrosis, non-enhancing tumor and edema) were also provided for the training images in .mha format. Additional sub-tasks included longitudinal evaluation of the segmentations for patients who had imaging from multiple time points. Finally, the third subtask was to classify the tumors into one of the three classes (Low Grade II, Low Grade III, and High Grade IV glioblastoma multiforme (GBM)). However, sub-tasks 2 and 3 were primarily pushed out to future years.
...
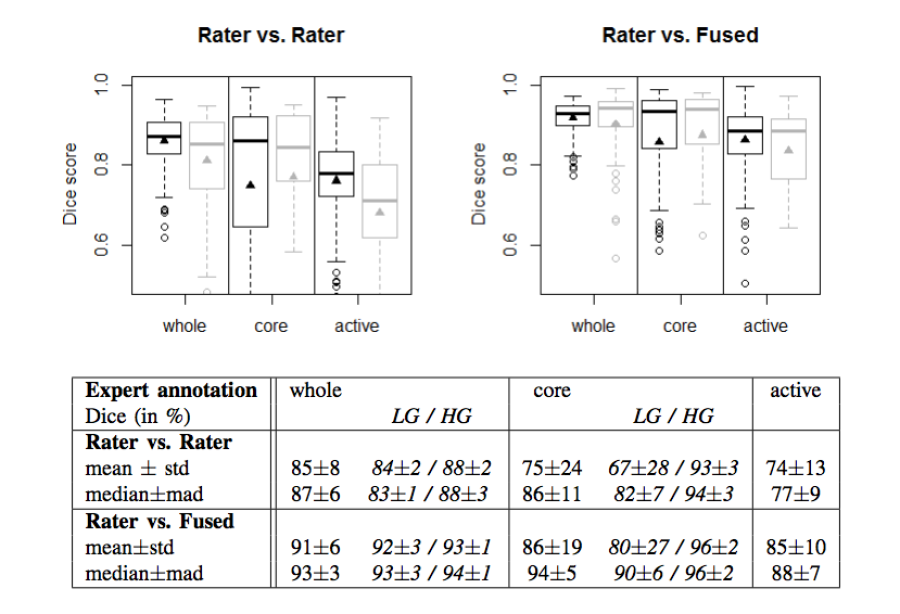
The MICCAI-BraTS challenge highlighted a number of findings that mirrored experiences from other domains. These
- The agreement between experts in is not perfect (~0.8 Dice score).
- The agreement (between experts and between algorithms) is highest for the whole tumor and relatively poor for areas of necrosis and non-enhancing tumor.
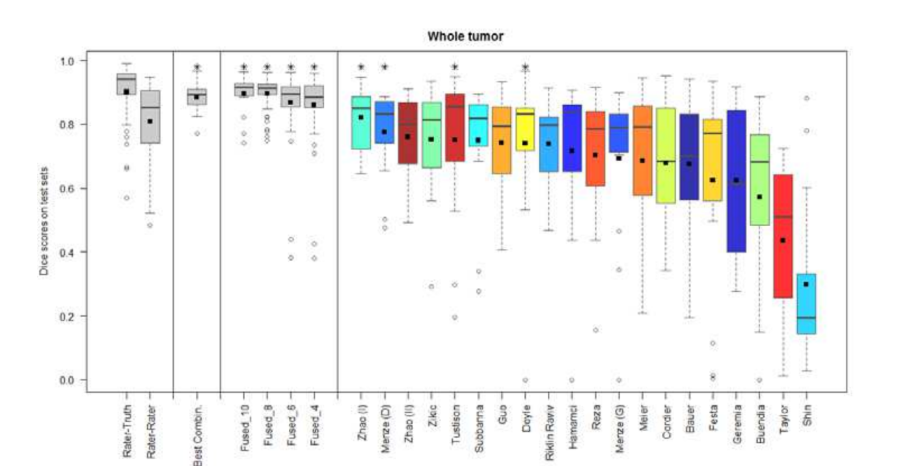
- Combining segmentations created by "best" algorithms created a segmentation that achieves overlap with consensus "expert" labels that approaches inter-rater overlap.
- This approach can be used to automatically create large labeled datasets.
- However, there are cases where this does not work and we still need to validate a subset of images with human experts.
Figure 2. Dice coefficients of inter-rater agreement and of rater vs. fused label maps
Figure 3. Dice coefficients of individual algorithms and fused results indicating improvement with label fusion
...
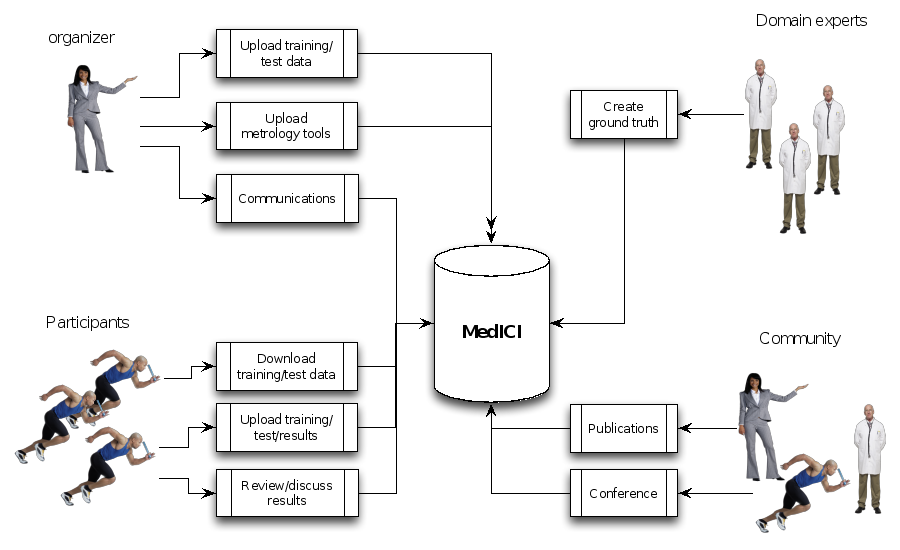
Below is a workflow diagram that describes the various stakeholders in the challenge and their tasks.
Figure 4. Challenge stakeholders and their tasks
...
These platforms typically charge a hosting fee and offering monetary rewards is pretty common. They have large communities (hundreds of thousands) of registered users and coders and can be a way to introduce the problem to communities outside the core domain expert academic researchers and get solutions that are novel in the domain.
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
The metrics that Kaggle supports
| multiexcerpt-include | ||||||
|---|---|---|---|---|---|---|
|
Error Metrics for Regression Problems
...
- Mean F Score
- Mean Consequential Error
- Mean Average Precision@nPrecision
- Multi Class Log Loss
- Hamming Loss
- Mean Utility
...
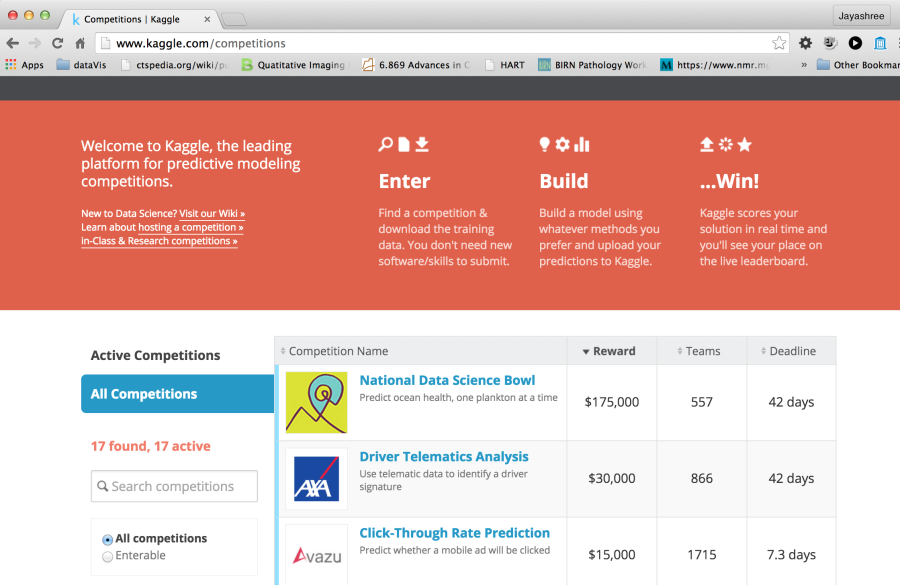
Figure 5. Portal for Kaggle, a leading website for challenges for data scientists
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
Challenge Post has been used to organize hackathons,
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
Open Source
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
...
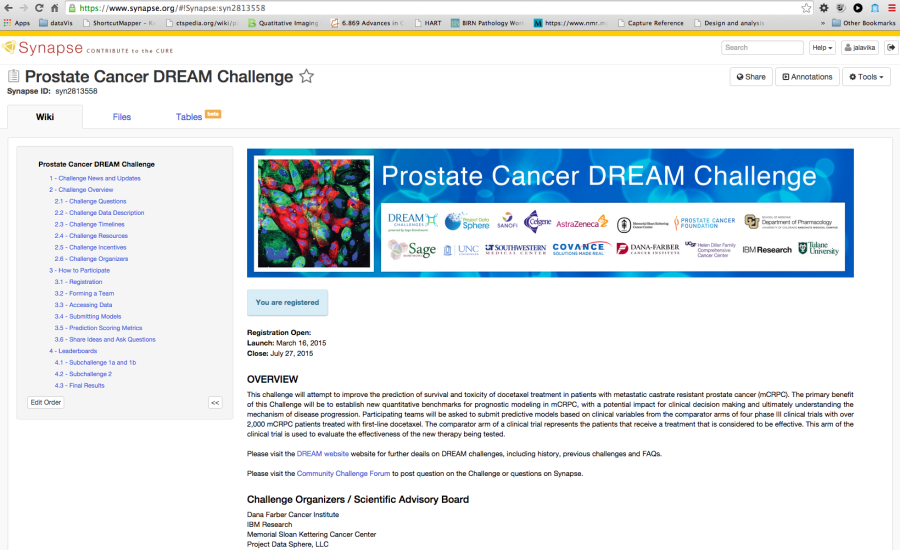
Figure 7. Example Challenge hosted in Synapse
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
The main steps to create a new challenge are:
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
- Create a project
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Add pages
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Making
- Create a project
- Add pages
- Making uploaded files available for download
- Allowing others to register for your project
- Make your project appear in the projects overview
- Allow file uploads
- Including content from files on a page
- Allow others to edit the project
- Changing colors and other styling
- Project data folder
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Allowing others to register for your project
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Make your project appear in the projects overview
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Allow file uploads
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Including content from files on a page
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Allow others to edit the project
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Changing colors and other styling
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Project data folder
Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include - Page permissions
Page permissionsMultiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to Include
However, at this time, there is limited support for automatic evaluation of submitted results, results presentation, native support for medical images although many of these features are planned.
The HubZero
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
...
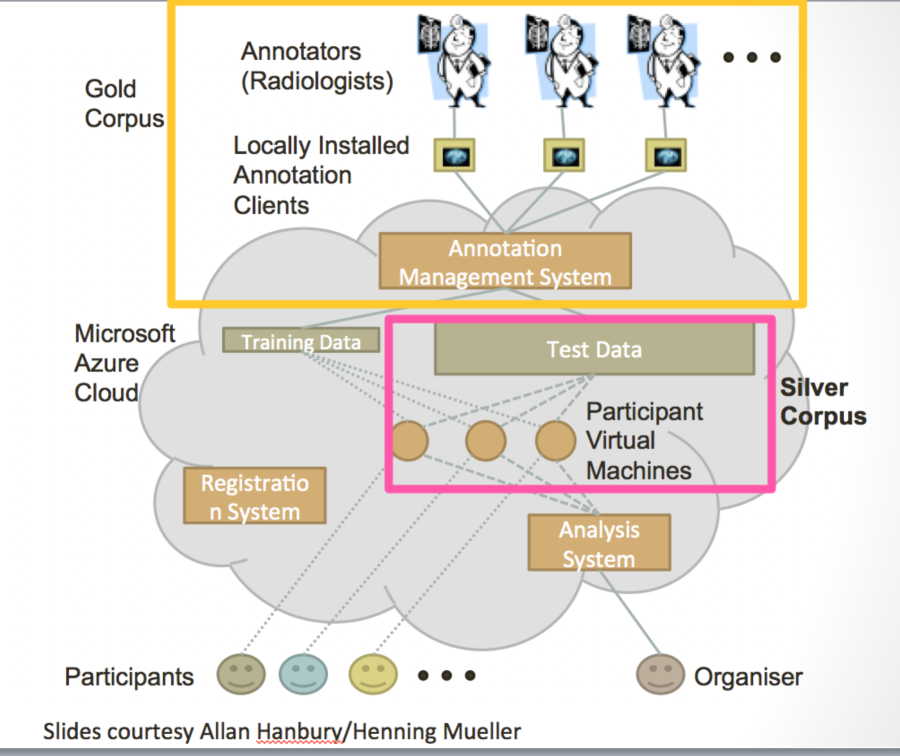
Figure 8. Schematic diagrams of the VISCERAL system for cloud-based challenges
The MIDAS platform has been used to host a couple of imaging challenges. A special been used to host a couple of imaging challenges. A special
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
...
Below is a table that rates the relative merits of the most relevant frameworks that we evaluated. The scale is 1-5 where 1 indicates excellent support for the feature while 5 indicates that that feature is not currently part of the system or there is limited support. We have included Kaggle as a representative "paid" framework for comparison. The other five frameworks are the open-source frameworks that were seriously considered in this comparison.
| Kaggle | Synapse | HubZero (challenges/projects) | COMIC | VISCERAL | CodaLab | |
|---|---|---|---|---|---|---|
| Ease of setting up new challenge | 2/4 (if new metrics need to be used) | 2 | 2/5 | 2 | 3 | 1 |
Cost (own server/hosting options) | $10-$25k/challenge | Free/hosted | Free/hosted | Free/hosted | Free/Azure costs | Free/hosted |
License | Commercial | OS | OS | OS | OS | OS |
Ease of extensibility | 5 | 4 | 4 | 2 | 3 | 2 |
Cloud support for algorithms | 4 | 3 | 3 | 4 | 1 | 3 |
Maturity | 1 | 1 | 1/5 | 3 | 4 | 3 |
Flexibility |
|
|
|
|
|
|
Number of users | 1 | 1 | 1/5 | 3 | 3 | 3 |
Types of challenges | 1 | 1 | 1 | 3 | 1 | 1 |
Native imaging support | No | No | No | Yes | Limited | No |
API to access data, code | 5 | 1 | 3 | 4 | 4 | 4 |
...
The web portal is the single point of entry for the participants. Historically, this would have information about the challenge, potentially host the data and provide a submission site for the user to upload results. The challenge organizer could also provide the results of the challenge at this page. Many challenges have wikis and announcement pages as well as forums. A good example of active discussion forums can be found at the Kaggle
| Multiexcerpt include | ||||||
|---|---|---|---|---|---|---|
|
...
Although Kaggle, Innocentive and Topcoder all have platforms that have been used extensively for a really wide range of challenges, these were excluded from further consideration (and from the above table) as the platforms since they are not open source and cannot be modified.
...