To Print the Guide
We recommend you print one wiki page of the guide at a time. To do this, click the printer icon at the top right of the page; then from the browser File menu, choose Print. Printing multiple pages at one time is more complex. For instructions, refer to How do I print multiple pages?.
Searching for Compounds Associated with a Disease
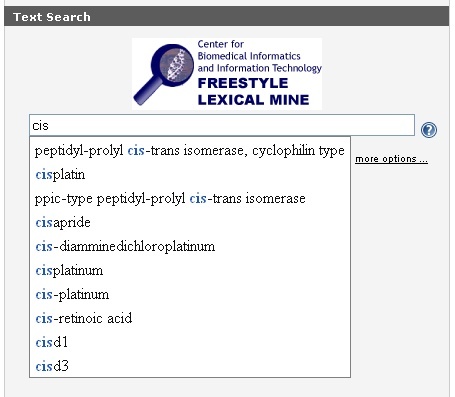
To use the caBIO Portal Freestyle Lexical Mine tool to find genes that are associated with a pharmacological substance, begin typing keywords into the search field. Because the tool suggests caBIO terms that match the characters you have entered, it is relatively easy to find a compound search term that matches your desired compound concept.
If you would like to search for any compound term that contains a string use the special character "*." For example, "cisplatin" would only return objects that are associated with this exact compound term, but "cisp*" will retrieve objects that are associated with any term that contains the string "cisp."
After entering a search term (1), click the Submit button (2) to retrieve results. Although you may limit your search by clicking on the more options ... link, this is not required (3).
Tip
If you have cannot find an appropriate search term, click the Contact Us link for help.
Search Results
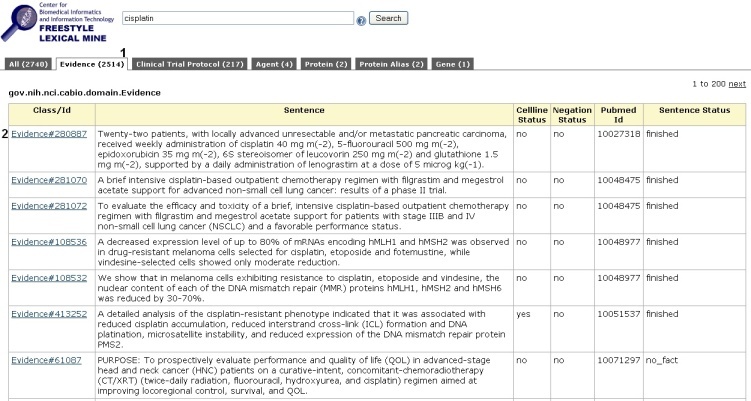
The caBIO Portal Freestyle Lexical Mine will retrieve objects with attributes that match your search term. These objects are grouped by type, which are shown as tabs at the top of the results page. To view genes that are associated with your compound term, click the Evidence tab at the top of the page (1).
Each row in the Evidence results table is a truncated view of an Evidence object (i.e., not all attributes and methods are shown on this page), where the columns include:
- the class and identifier (
Class/Id), - evidence of the gene-compound association (
Sentence), - whether the evidence was collected from experiments involving cell lines (
Cellline Status), - whether the evidence is negative (i.e., gene X is not associated with compound Y;
Negation Status), - the PubMed identifier for the abstract from which the evidence was extracted (
Pubmed Id), and - whether the status of the sentence (
Sentence Status).
For additional information on these attributes of the Evidence type objects, refer to the section Data, Metadata, and Annotations.
Note
Although the Cancer Gene Index refers to pharmacological substances as "compounds" or "drugs," caBIO and the NCI Thesaurus use the term "agent" for the same concept.
Finding the Gene Associated with the Evidence
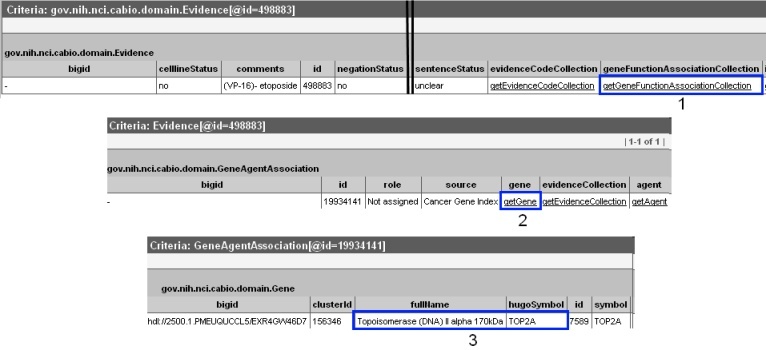
To discover which gene is associated with each piece of evidence, click on the Class/Id link for the desired object. This will open the full Evidence type object. Scroll over to the right, and click on getGeneFunctionAssociationCollection method link (1) to view the Gene Agent Association type object. This object has a role attribute that contains one or more Role Codes or Role Details that describe the nature of the gene-compound/agent relationship, as well as a notation that the Cancer Gene Index is the source of these data.
Warning
If you do not want to spend time navigating through the caBIO object model for candidate gene-compound associations that were found to be false positives, select only Evidence objects where the Sentence Status is finished and Negation Indicator} is {{no.
Note
A single piece of evidence may have multiple Role Codes and Role Details describing the gene-compound association, and the evidence may also describe gene-disease associations. Thus, after clicking the getGeneFunctionAssociationCollection link, you may see multiple retrieved objects of type Gene Agent Association, gov.nih.nci.cabio.domain.GeneAgentAssociation, and even multiple object retrieved records of type Gene Disease Association, gov.nih.nci.cabio.domain.GeneDiseaseAssociation.
Click on the getGene link (2) to access the related Gene object (bottom panel). This Gene object contains the full name and HUGO Gene Symbol in the fullName and hugoSymbol columns (3), for example, for the gene associated with the compound of interest and a specific piece of evidence.
To explore additional genes associated with the compound term, navigate back to the evidence page and repeat this process.
Be Careful
If you find yourself in a part of the object model that you do not understand or if you get confused, stop and navigate your web browser back to the search results page with the Evidence tab.
For your reference, the subset of caBIO classes that are related to the Cancer Gene Index are shown here. The full caBIO model is available on the caBIO gForge page, but you must have the Enterprise Architect modeling tool to view this file.
Agent Ontologies
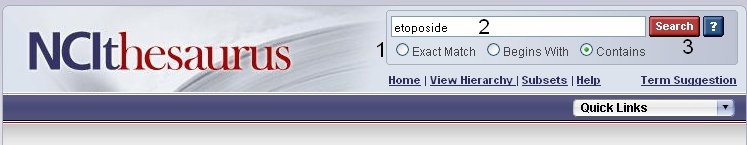
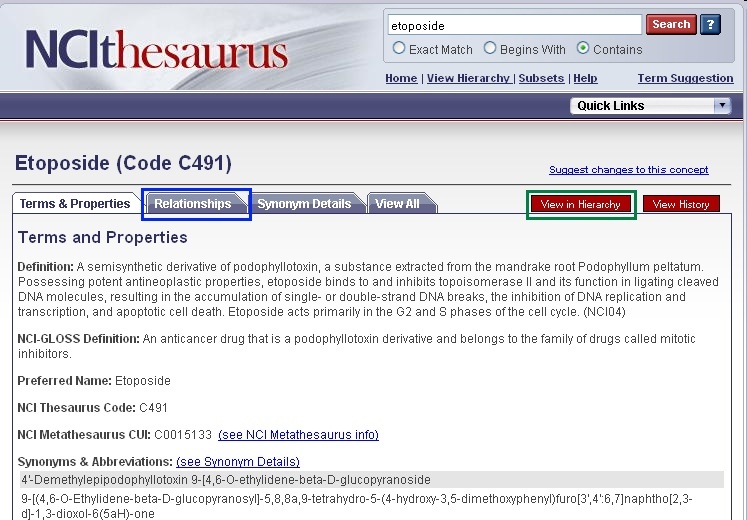
If you would like to search for genes that are associated with parent, sister, or child concepts to your compound/agent search term, you must use the NCI Thesaurus. To view these ontologies in the NCI Thesaurus, open a new browser tab or window and navigate to the NCI Thesaurus web page, enter in your compound term (2, "etoposide") or NCI Thesaurus concept code (e.g., "C491"), and click the Search button (3). If required, select your exact search term from the list to view the NCI Term page
Identifier Tip
The EVS Identifier for a term is also its NCI Thesaurus Concept Code.
You may view parent and child terms for any compound term by clicking on the Relationships tab (blue box). For example, "etoposide" has the no child terms and the parent term " Epipodophyllotoxin Compound." Alternatively, if you would like to view parent, sister, and child concepts as well as where your term fits in the entire agent hierarchy, click the red View in Hierarchy button (green box).