This chapter describes the processes for creating and managing studies in caIntegrator. Topics in this chapter include:
You can create a caIntegrator study by importing subject annotation study data, genomics data and imaging data You can incorporate a combination of spreadsheets/files and existing caGrid applications as source data. Each instance of caIntegrator can support multiple studies. As the manager creating a study, it is important that you understand the study well and that the data you wish to aggregate has been submitted to the applications whose data can be integrated in caIntegrator.
As you create the study, you define its structure, identifying the data sources and mapping the data between different source data. After the study has been created and deployed, you can perform analyses of the data in the study.
Only a user with a Study Manager role can create a study. For more information, see caIntegrator Roles. |
When you create a study, you must specify different data-types (subject annotation, array, image, etc), data sources (caGrid applications – caArray and NBIA) and map the data, (patient to sample, image series, etc.).
To create a new study, follow these steps:
The Edit Study page, as shown in the following figure, displays the Name and Description that you entered for a new study, or for an existing study that you are editing.
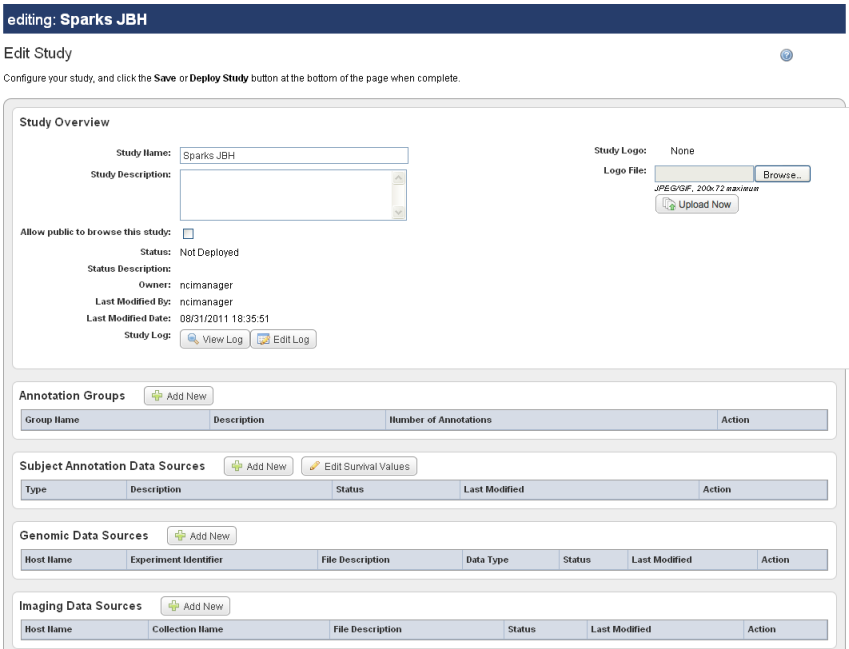
To continue creating a study or to modify a study, complete these steps:
You can save the study at any point in the process of creating it. You can resume the definition and deployment process later. |
To continue creating the study, you can add subject annotation data sources, genomic data sources or imaging data sources.
On the Edit Study page, as a study manager you can open a detailed log for the study.
See also #Study Log.
One of the most important tasks in creating a study in caIntegrator is in properly annotating the data. Each annotation has a definition you must identify. Because the process can be quite complex, you might want to review the following steps for working with annotations.
Annotation Workflow Summary:
This topic opens from both the Create Annotation Group page and the Edit Annotation Group page. If you plan to create a group, continue with this topic. If you plan to edit an annotation group, see #Editing an Annotation Group.
An annotation group is a group of annotation definitions configured in a CSV file. This feature is primarily meant for the Study Manager who knows that they have tightly restricted vocabulary definitions that are relevant to a study. In this optional step, you can review the uploaded Group Definition Source file before assigning the appropriate definitions for your study.
To add an annotation group, follow these steps:
When you open the Define Fields for Subject Data page, the annotation definitions in the file you uploaded display on the page, available for assignment in the study. Additionally, you can view the definitions by viewing the annotation group listed in the first column of the matrix.
Annotation definitions by default are visible only to the Study Manager's group. They are not visible to all caIntegrator users, unless you change the visibility for each. |
This topic opens from the Edit Annotation Group page. You may want to refer to #Adding an Annotation Group if you are adding a group for the first time.
To edit an annotation group, on the Edit Study page for a study with an existing annotation group, click the Edit Group button.
The Edit Study page, described in #Creating/Editing a Study, opens after you save a new study or click to edit an existing study.
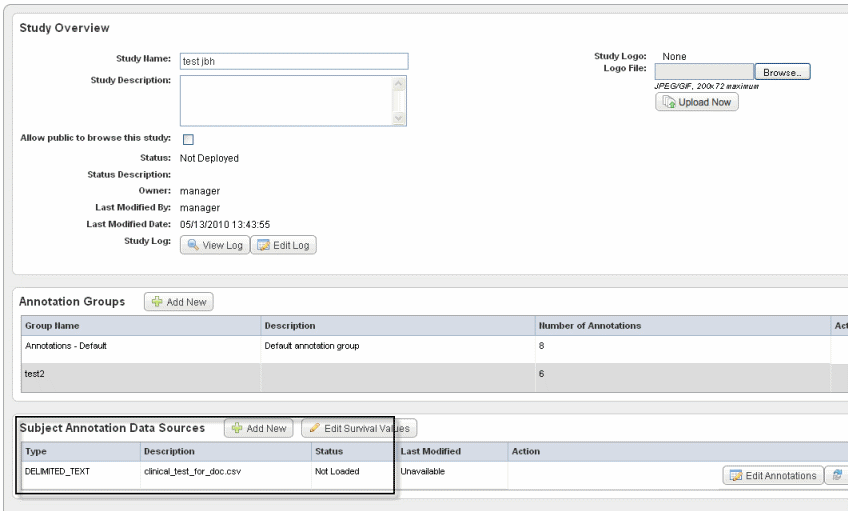
To add subject annotation metadata on this page, follow these steps:
After the data file is uploaded to this study, it will be listed in the Subject Annotation Data Sources section of the Edit Study page.
From this page you can initiate editing the annotations. In the Subject Annotation Data Sources section, click Edit Annotations corresponding to the subject annotations that have been uploaded for the study. This open the #Define Fields Page for Editing Annotations.
The Define Fields for Subject Data page, shown in the following figure, opens when you click Edit Annotations in the Subject Annotation Data Sources or the Image Data Sources section of the Edit Study page. The exception to this is if you have not yet imported annotations for the imaging data for the study, In that case, when you click the Edit Annotations button in the Imaging Data Sources section, a page opens where you can identify and upload image annotation data. See #Adding or Editing Image Annotations.
If this Define Fields page opens after clicking the Edit Annotations button, working with this page is identical for both subject and image annotations
The MOST important steps in creating annotation definitions on this page are these:
The first column of the table on this page displays annotation groups that have been created for this study. For more information, see #Adding an Annotation Group.
To add subject or image annotation metadata in this page, follow these steps:
When you click Change Assignment on the Define Fields page, the Assign Annotation Definition for Field Descriptor dialog box opens. On this page you can change the column type and the field definition for the specific data field you selected.
When you change an assignment, you must make sure the data types match--numeric, etc. |
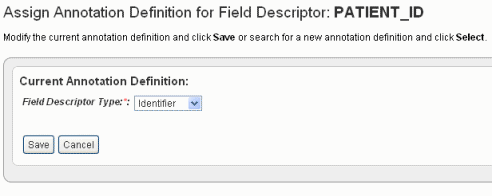
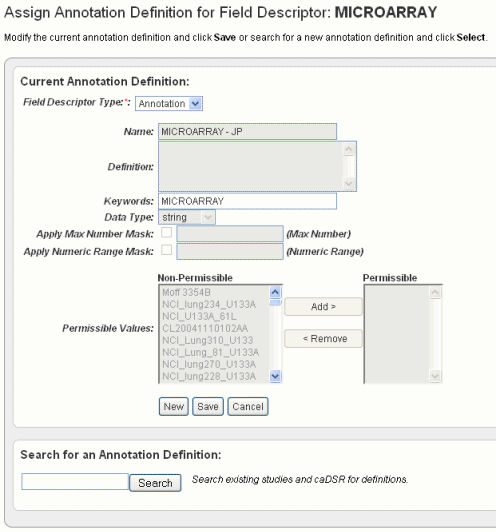
Annotation Field |
Field Description |
|---|---|
Name |
Enter the name for the annotation. |
Definition |
Enter the term(s) that define the annotation. |
Keywords |
Insert keyword(s) that could be used to find the annotation in a search, separated by commas. |
Data Type |
Select a string (default), numeric, or date. |
Apply Max Number Mask |
This field is available only for numeric-type annotations, or when a new definition is created. This feature is unavailable when permissible values are present. |
Apply Numeric Range Mask |
This field is available only for numeric-type annotations, or when a new definition is created. This feature is unavailable when permissible values are present. |
Permissible/Non-permissible Values |
Note: The first time you load a file, before you assign annotation definitions, step #3 in #Assigning An Identifier or Annotation, these panels may be blank. If the column header for the data is already "recognizable" by caIntegrator, the system makes a "guess" about the data type and assigns the values to the data type in the newly uploaded file. They will display in the Non-permissible values sections initially. Use the Add and Remove buttons to move the values shown from one list to the other, as appropriate. |
When you select or change annotation definitions by selecting matching definitions (described in #Searching for Annotation Definitions), this may add (or change) the list of non-permissible values in this section.
If you leave all values for a field in the Non-permissible panel, then when you do a study search, you can enter free text in the query criteria for this field.
If there are items in the Permissible values list, then the values for this annotation are restricted to only those values. When you perform a study search, you will select from a list of these values when querying this field. If there are no items in the permissible values list then the field is considered free to contain any value.
To edit a field's permissible values, you must change the annotation definition. You can do this even after a study has been deployed.
Note: You cannot edit permissible values in an existing annotation definition. To change permissible values, you must create a new annotation.
An alternative to creating a new definition is to search for annotation definitions already present in caIntegrator studies or in caDSR.
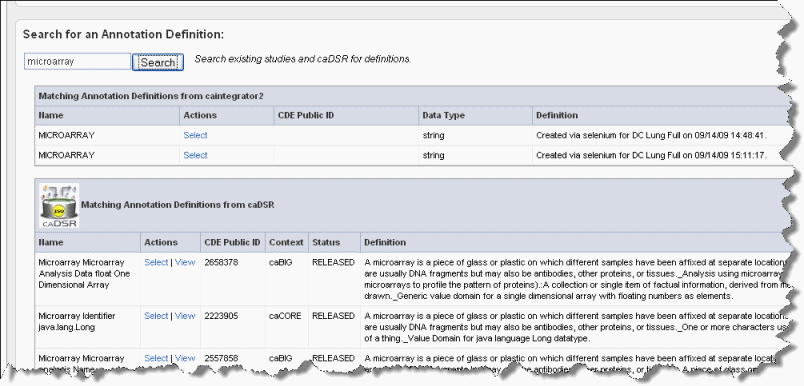
To view the definitions corresponding to any of the "Matching Annotation Definitions", which are those currently found in other caIntegrator studies, click the \[term\], such as "age", hypertext link. The definition then appears in the Current Annotation Definition segment of the page just above. |
When you click the link for a definition, that assigns the definition to the Define Fields for Subject Data page, and it also closes the Annotation Definition page. You can modify any portion of the definition, as described in Step 6 in #Assigning an Identifier or Annotation. |
Take care before you add a caDSR definition that it says exactly what you want. caDSR definitions can have minor nuances that require specific and limited applications of their use. |
If you have not clicked Select for alternate definitions in this dialog box, then click Save to return to the Define Field...dialog box without making any definition changes. |
Saving your entries in this way saves the study by name and description, but does not deploy the study. See #Deploying the Study. |
You can add as many files as are necessary for a study. Patients 1-20 in first file, 21-40 in second file, or many patients in first file and annotations in second file, etc. As long as IDs are defined correctly, it works. |
You can change assignments even after the study is deployed, using the Edit feature. For more information, see #Creating/Editing a Study. |
The Manage Studies page opens when the study is deployed. The Deployed status is indicated on the Manage Studies page as well as the Edit Study page. For more information, see #Managing a Study.
You can continue to perform other tasks in caIntegrator while deployment is in process.
See also #Deploying the Study.
You can repeatedly upload additional or updated subject annotations, samples, image data, array data to the study at later intervals. These later imports do not remove any existing data; they instead insert any new subjects or update annotations for existing subjects. |
Survival value is the length of time a patient lived. If you plan to analyze your caIntegrator data to create a Kaplan-Meier (K-M) Plot, then during the Annotation Definition process described above in #Assigning an Identifier or Annotation, you should do one of two things:
Setting survival values is optional if you do not plan to use the K-M plot analysis feature or if you do not have this kind of data (survival values) in the file. |
In caIntegrator, survival values are not pre-defined in the databases when you load the data. However, you can review and define survival value ranges in a data set you are uploading to a study. To be able to do so, you need to understand the kind of data that can comprise the survival values.
To set up survival values, follow these steps:
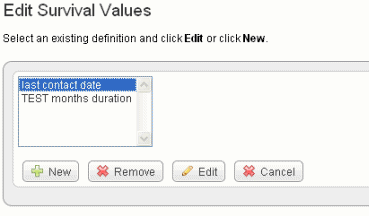
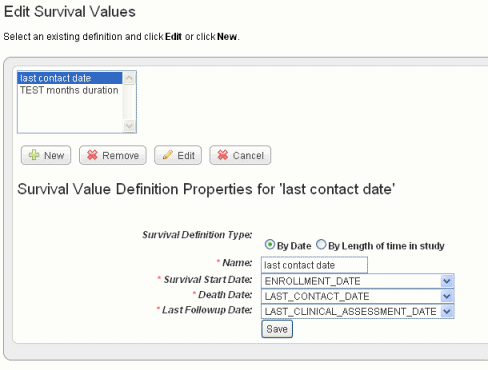
Field Type |
Description |
|---|---|
Survival Definition Type |
Select whether the survival time is defined by dates or length of time subject was in the study. |
Name |
Enter a unique name that adequately describes the survival values you are defining here. Example: Survival from Enrollment Date or Survival from Treatment Start. The name you enter displays later when you are selecting survivals to create the K-M plot. |
Survival Length Units |
Select the appropriate units for this data. |
Survival Start Date |
Select the column header for this data. |
Death Date |
Select the column header for this data. |
Last Followup Date |
Select the column header for this data. |
For data analysis using survival values, see Creating Kaplan-Meier Plots.
Genomic data that is parsed and stored in caArray can be analyzed in caIntegrator. Additionally, supplemental files in caArray that have not been parsed can be uploaded and analyzed in caIntegrator. |
Once you have loaded subject annotation data and identified subject IDs, you can add one or more sets of array genomic sample data from caArray to the study. caIntegrator maps the data by sample IDs to the subject IDs in the subject annotation data, covered in this section, or you can load imaging files from NBIA, also mapped by IDs to the subject data. This is discussed in #Working with Imaging Data. You can also edit genomic data information that you have already added to the study. Genomic sample data and imaging data are independent of each other, so neither is required before loading the other.
It is essential that you are well acquainted with the data you are working with--the subject annotation data, and the corresponding array data in caArray.
caIntegrator supports a limited number of array platforms. For more information, see #Managing Platforms.
To add genomic data to your caIntegrator study, follow these steps:
Field Types |
Field Description |
|
|---|---|---|
caArray Web URL |
Enter the URL for the caArray instance to be used for the genomic data sources. This will enable a user to link to the referenced caArray experiment from the study summary page. |
|
caArray Host Name |
Enter the hostname for your local installation or for the CBIIT installation of caArray. If you misspell it, you will receive an error message. |
|
caArray JNDI Port |
Enter the appropriate server port. See your administrator for more information. Example: For the CBIIT installation of caArray, enter 8080. |
|
caArray Username and caArray Password |
If the data is private, you must enter your caArray account user name and password; you must have permissions in caArray for the experiment. If the data is public, you can leave these fields blank. |
|
caArray Experiment ID |
Enter the caArray Experiment ID which you know corresponds with the subject annotation data you uploaded. Example: Public experiment "beer-00196" on the CBIIT installation of caArray (array.nci.nih.gov). If you misspell your entry, you will receive an error message. |
|
Vendor |
Select either Agilent or Affymetrix. |
|
Data Type |
Select Expression or Copy Number. |
|
Platform |
If appropriate, select the Agilent or Affymetrix platform.
|
|
Central Tendency for Technical Replicates |
If more than one hybridization is found for the reporter, the hybridizations will be represented by this method. |
|
Indicate if technical replicates have high statistical variability |
If more than one hybridization is found, checking this box will display a ** in the genomic search results when a reporter value has high statistical variability. |
|
Standard Deviation Type |
When the checkbox for indicating if technical replicates have high statistical variability is checked, this parameter becomes available. Select in the drop-down the calculation to be used to determine whether or not to display a ** (see previous bullet point). |
|
Standard Deviation Threshold |
When the checkbox for indicating if technical replicates have high statistical variability is checked, this parameter becomes available. This is the threshold at which the Standard Deviation Type is exceeded and the reporter is marked with a **. |
caIntegrator goes to caArray, validates the information you have entered here, finds the experiment and retrieves all the sample IDs in the experiment. Once this finishes, the experiment information displays on the caIntegrator Edit Study page under the Genomic Data Sources section, as shown in the following figure.
If you want to redefine the caArray experiment information, you can edit it. Click the Edit link corresponding to the Experiment ID. The Edit Genomic Data Source dialog box reopens, allowing you to edit the information. |
Because the goal of caIntegrator is to integrate data from subject annotation, genomic and imaging data sources, data from uploaded source files must be mapped to each other. Mapping files can map to caArray genomic data of two types: "imported and parsed" and that stored in supplemental files.
You, as the caIntegrator study manager, must create a Subject to Sample mapping file and then import it into caIntegrator before following the actual mapping steps. This file provides caIntegrator with the information for mapping patients to caArray samples.
Only one of the last 2 columns is used: a single sample per file uses the Value Header column; multiple samples per file used Sample Header column. Unused columns are blank. |
The following figure shows an example multiple sample mapping file in CSV format.
Supplemental files from caArray for mapping data must be configured appropriately. For information, see Supplemental Files Configuration. |
To map the samples from the caArray experiment to the subjects in the subject annotation data you uploaded, follow these steps:

Field |
Description |
|---|---|
caArray Host Name |
Enter the hostname for your local installation or for the CBIIT installation of caArray. If you misspell it, you will receive an error message. |
caArray JNDI Port |
Enter the appropriate server port. See your administrator for more information. Example: For the CBIIT installation of caArray, enter 8080. |
caArray Username |
Enter your caArray account user name and password; you must have permissions in caArray for the experiment if it is private. If the data is public, you can leave this field blank. |
caArray Experiment ID |
Enter the caArray Experiment ID which you know corresponds with the subject annotation data you uploaded. Example: Public experiment "beer-00196" on the CBIIT installation of caArray (array.nci.nih.gov). If you misspell your entry, you will receive an error message. |

If you have already mapped samples, when you first open this page they are listed in the Samples Mapped to Subjects section. If you have not already mapped samples, all of the samples in the caArray experiment you selected are listed as unmapped, because caIntegrator does not know how these sample names correlate to the patient data in the subject annotation file until you upload the subject to sample mapping file. |
A Control Samples file is used to calculate fold change data, which compares "tumor" sample gene expression in the caArray experiment to the control samples to identify those that exhibit up or down gene regulation. Control samples can be the "normal" samples, but that is not always the case.
To upload the control samples, follow these steps:
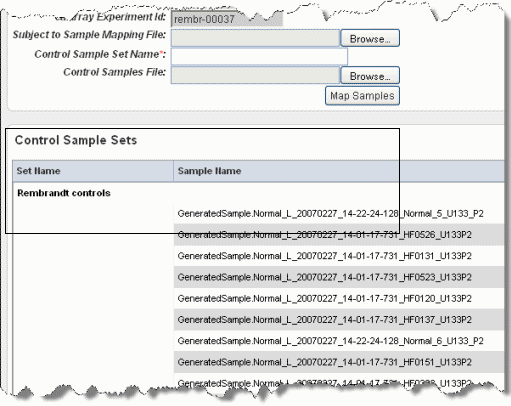
This information will be used when performing other tasks in caIntegrator, to be described in other sections.
If a Control Set is to be used in Gene Expression For Annotation, or Gene Expression Plots for Annotation Query, then the control set should be composed of only samples which are mapped to subjects. |
You can add copy number data for a genomic data source by uploading the mapping file. This allows you to configure parameters to be used when segmentation data is being configured.
The name specified in the third column of the mapping file is specific for each array manufacturer:
To add copy number data relating to the genomic data you are adding, follow these steps:
This link is available only if you have uploaded copy number data and you are configuring a Copy Number data type (as indicated by the Data Type column on the Edit Study page). |
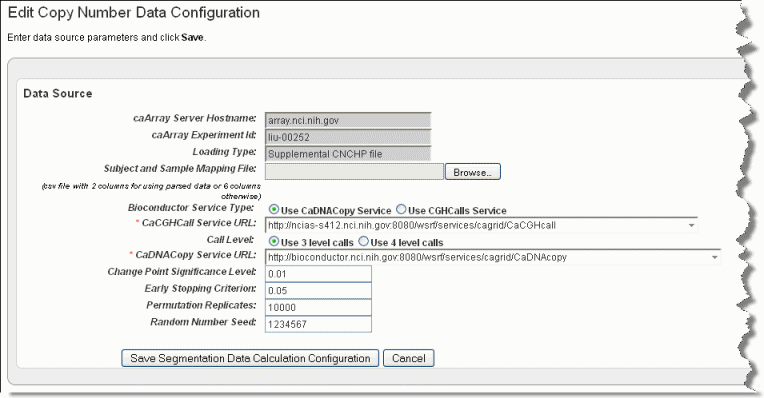
Field |
Description |
|
|---|---|---|
caArray Service Host Name |
Enter the hostname for your local installation or for the CBIIT installation of caArray. If you misspell it, you will receive an error message. |
|
caArray Experiment ID |
Enter the caArray Experiment ID which you know corresponds with the copy number data. |
|
Loading Type |
Enter the Loading Type of the data file you plan to map. |
|
Subject and Sample Mapping File |
Browse for the appropriate CN mapping file. The file must be a CSV file with 3 column format for mapping data files (format: subject id, sample id, file name). Supplemental data uses 6 column-files. |
|
Bioconductor Service Type |
This is the type of bioconductor module that will be used for segmentation. Select between the two options: DNAcopy or CGHcall. |
|
caCGHcall Service URL |
Enter the URL for the grid segmentation service used to access the caCGHcall service. For more information, see CGHcall |
|
Call Level |
An input parameter to CGHcall. This is the number of discrete values used to represent the copy number level. Select between two options: 3 (consisting of discrete values of -1, 0, 1) or 4 (consisting of discrete values -1, 0, 1, 2) |
|
caDNACopy Service URL |
Control for selecting the URL which hosts the caDNACopy grid service. For more information, see DNAcopy |
. |
Change Point Significance Level |
Significance levels for the test to accept change-points |
|
Early Stopping Criterion |
The sequential boundary used to stop and declare a change |
|
Permutation Replicates |
The number of permutations used for p-value computation |
|
Random Number Seed |
The segmentation procedure uses a permutation reference distribution. This should be used if you plan to reproduce the results. |
After a study has been deployed and the genomic source has been loaded, you cannot change these copy number parameters without reloading the data from caArray first. |
Occasionally you may need to remap copy number data in a deployed study. To do so, follow these steps:
Once you have loaded subject annotation data and identified patient IDs, you can add either array genomic sample data from caArray which caIntegrator maps by sample IDs to the patient IDs in the subject annotation data, or you can upload image data from NBIA, also mapped by IDs to the subject data. Once you have configured an NBIA image data source for adding images, then you can import image annotation data for the images. Genomic sample data and imaging data are independent of each other, so neither is required before loading the other.
It is essential that you are well acquainted with the data you are working with--the subject annotation data, and the corresponding imaging data in NBIA.
To add images from NBIA to the study you are creating, follow these steps:
If you have already provided an imaging data source, it is listed in this section of the Edit Study page. To edit the imaging data source, click the Edit button which opens the same dialog box described in the following steps. |
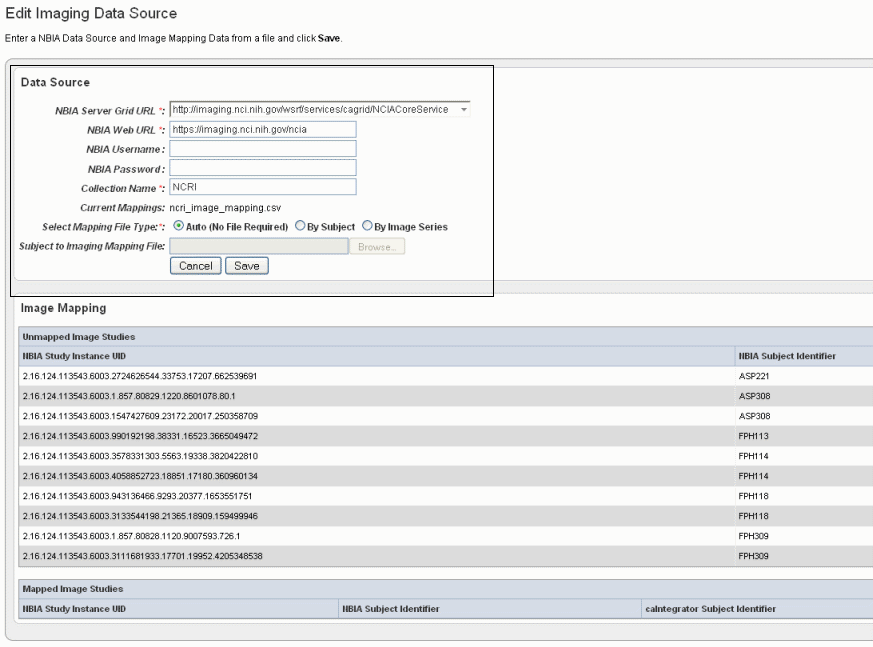
Fields |
Description |
|---|---|
NBIA Server Grid URL* |
Enter the URL for the grid connection to NBIA. |
NBIA Web URL |
Enter the URL of the web interface of the NBIA installation. |
NBIA Username and NBIA Password |
This information is not required, as currently all data in the NBIA grid is Public data. |
Collection Name |
Enter the name/source for the collection you want to retrieve. |
Current Mapping |
If a mapping file has already been uploaded to the study to map imaging data, the file name displays here. |
Select Mapping File Type |
Click to select the file type: |
Subject to Imaging Mapping File |
Click Browse to navigate to the appropriate subject annotation to imaging mapping file. See the Select Mapping File Type* field description. |
If mapping files have already been uploaded for the data sources you are editing, the Image Mapping tables of the dialog box show the mapping from NBIA Image Series Identifier to caIntegrator Subject Identifier. |
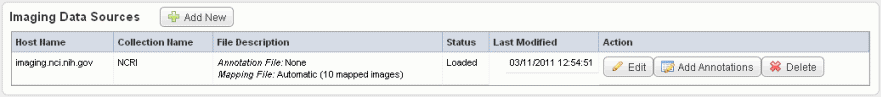
After you have configured an image data source with an NBIA Grid service and uploaded the image data, described in #Adding or Editing Image Data Files from NBIA, you can load image annotations into caIntegrator from a file in CSV format or through an Annotations and Image Markup (AIM) service.
The image data shown in the Imaging Data Sources section indicate whether or not annotations have already been imported from a file for these sources. See the marked area in the following figure. |
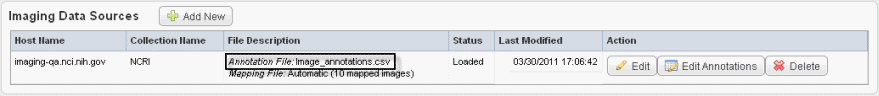
To add image annotations from a file, follow these steps:
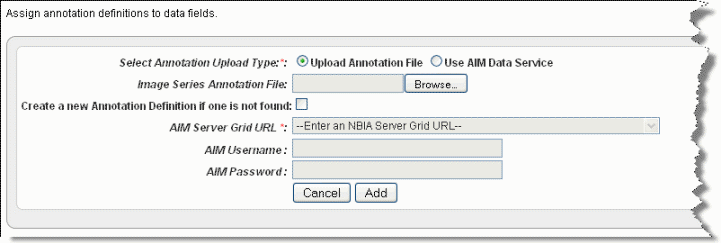
If you have not yet imported annotations, clicking this button opens the page from which you can import image annotations, shown in the following figure. Continue with the steps in this section. If you are editing annotations, clicking this button opens the Define Fields for Image Annotations dialog box where you can edit annotations; see #Define Fields Page for Editing Annotations. |
An image annotation CSV file must include an Image Series ID column. See the highlighted column in the following figure. |
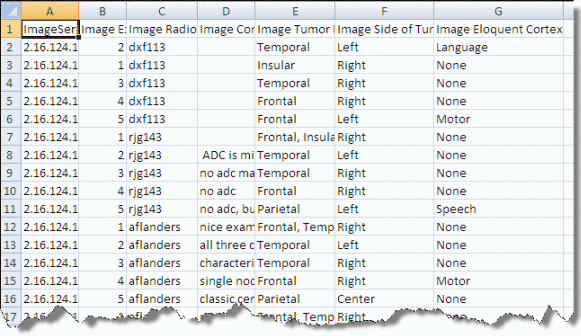
To load image annotations through an AIM service, follow these steps:
Using either method, the image annotations are uploaded to caIntegrator. After this occurs, when you click the Edit Annotations button, the system opens to the Define Fields for Imaging Data page where you can edit the annotations. This is the same page (with a customized title) as that described in #Define Fields Page for Editing Annotations. You must assign identifiers and annotations to the data in the same way you did with the subject annotation data. For more information, see #Assigning an Identifier or Annotation and #Searching for Annotation Definitions.
If you are a study manager, this feature on the Edit Study page allows you to configure a CSV file with URLs to be used as external links relevant to the study. This allows you to easily share or configure references.
To add an external link, follow these steps:
Once you have created external links for a study, when the study is open, an External Links section on the left sidebar of the page shows the link(s). An example is identified in the following figure.
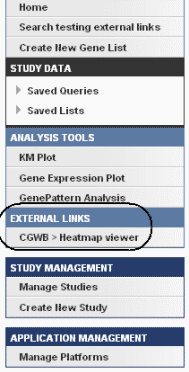
Click an external link to open a page that displays appropriately formatted web page links; an example is shown in the following figure.

When you are ready to deploy the study, click the Deploy Study button on the Edit Study page. caIntegrator retrieves the selected data from the data service(s) you defined and makes the study available to a study manager or to anyone else who may want to analyze the study's data. Using the Manage Studies feature, you can then configure and share data queries and data lists with all investigators who access the study.
Note that you can continue to work in caIntegrator while the study is being deployed.
A user without management privileges has no access to this section of caIntegrator. |
Once you have started to create a study or have deployed it, you can update the study in the following ways:
To update, edit or delete a study, follow these steps:
caIntegrator supports a limited number of array platforms, all of which originate from Agilent or Affymetrix. While they do not represent all of the platforms supported by caArray, caIntegrator must have array definitions loaded for the platforms it supports, and be able to properly load the data from caArray and parse it.
You can create a study without genomic data, but you cannot add genomic data to a caIntegrator study without a corresponding supported array platform. If you add more than one set of genomic data to the study, you can specify more than one platform for the study.
On the Manage Platforms page, you can identify, add or remove supported platforms.
To manage platforms in caIntegrator, follow these steps:
Tab-delimited .txt or .tsv Agilent platform annotation files must contain the following column headers: ProbeId, GeneSymbol, GeneName and Accessions. |
The platform deployment can be time-consuming. If the platform takes more than 12 hours to deploy, caIntegrator displays a "timed out" message. At that point, you can delete the platform, even if it has not loaded to the system.
Platform loading can fail if the manufacturer's platform annotation file is missing data. |