|
Page History
...
In the Genomic Data Sources section of the Edit Study page, for the data you have already added, click Configure Copy Number Databutton.
Info title Uploaded copy number data? This link is available only if you have uploaded copy number data and you are configuring a Copy Number data type (as indicated by the Data Type column on the Edit Study page).
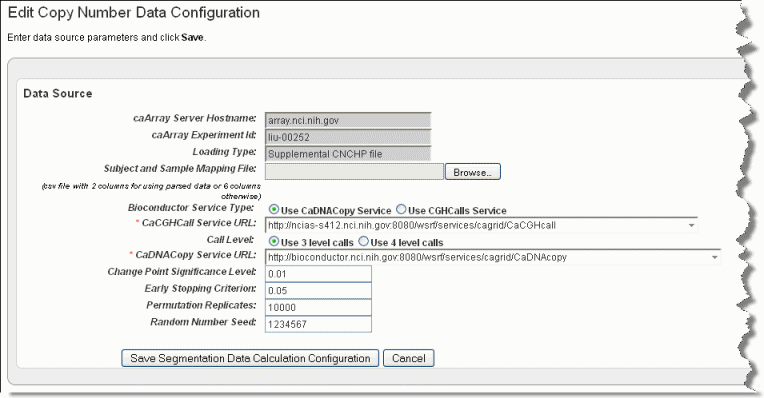
The Edit Copy Number page, shown in the following figure, opens.
Browse for and enter appropriate information to identify and retrieve the copy number mapping file. The fields are described in the following table. An asterisk indicates a required field.
Field
Description
caArray Service Host Name
Enter the hostname for your local installation or for the CBIIT installation of caArray. If you misspell it, you will receive an error message.
caArray Experiment ID
Enter the caArray Experiment ID which you know corresponds with the copy number data.
Loading Type
Enter the Loading Type of the data file you plan to map.
Subject and Sample Mapping File
Browse for the appropriate CN mapping file. The file must be a CSV file with 3 column format for mapping data files (format: subject id, sample id, file name). Supplemental data uses 6 column-files.
Bioconductor Service Type
This is the type of bioconductor module that will be used for segmentation. Select between the two options: DNAcopy or CGHcall.
caCGHcall Service URL
{multi-excerpt-include: Enter the URL for the grid segmentation service used to access the caCGHcall service. For more information, see CGHcall
Wiki Markup Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to
|name=ExitDisclaimer|nopanel=true}Include
.Call Level
An input parameter to CGHcall. This is the number of discrete values used to represent the copy number level. Select between two options: 3 (consisting of discrete values of -1, 0, 1) or 4 (consisting of discrete values -1, 0, 1, 2)
caDNACopy Service URL
{multi-excerpt-include: Control for selecting the URL which hosts the caDNACopy grid service. For more information, see DNAcopy
Wiki Markup Multiexcerpt include nopanel true MultiExcerptName ExitDisclaimer PageWithExcerpt wikicontent:Exit Disclaimer to
|name=ExitDisclaimer.|nopanel=true}Include .
Change Point Significance Level
Significance levels for the test to accept change-points
Early Stopping Criterion
The sequential boundary used to stop and declare a change
Permutation Replicates
The number of permutations used for p-value computation
Random Number Seed
The segmentation procedure uses a permutation reference distribution. This should be used if you plan to reproduce the results.
Click Save Segmentation Data Calculation Configurationfor a genomic data source. On the screen upload a copy number mapping file and configure the parameters to be sent when computing segmentation data.
Note title Be Careful After a study has been deployed and the genomic source has been loaded, you cannot change these copy number parameters without reloading the data from caArray first.
...