|
Page History
| Scrollbar | ||
|---|---|---|
|
| Section | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
You can use these documents to uncover fact-based associations between genes and diseases or genes and compounds. In addition, you can evaluate the evidence from which these associations were extracted (that is, the sentences from MEDLINE abstracts) and the codes and details that describe these associations. Other annotations, such as sentence status flags, negation indicators, cell line indicators, and organism data are information-rich and may be used to filter limit queries of the data, as requiredfor example. Additional information is available in the #Using the Cancer Gene Index subsection below. Cancer Gene Index DTDsUsing the Cancer Gene Index subsection of this document.
Cancer Gene Index DTDsDescriptions of In order to use the XML, you must first understand the DTD elements (which correspond to the XML elements) and how to interpret information within the elements. The Gene-Disease and Gene-Compound DTD and XML documents each contain thirty elements. Twenty-seven of these elements and the sequence of these elements are identical for the Gene-Disease and Gene-Compound documents. Only the three elements specifically referring to or containing data on diseases or compounds differ between the documentsDTD elements are provided below. Description of the Cancer Gene Index Gene-Disease DTD Elements
A Some of the genes in the gene-disease concept pairs are not included in the HGNC or LocusLink and, thus, the text contents for these elements will be "0" Description of the Cancer Gene Index Gene-Compound DTD Elements
A Some of the genes in the gene-compound concept pairs are not included in the HGNC or LocusLink and, thus, the text contents for these elements will be "0" Additional DTD InformationThe Gene-Disease and Gene-Compound DTD elements include meaningful parenthetical information and special characters. Consider the following example Cancer Gene Index DTD elementsexamples:
Cancer Gene Index elements not only contain child elements and text elements, but also information about the presence of child elements and the number of times a particular element can recur. Elements with one or more child elements declare the name(s) of the child elements as comma-separated lists inside parentheses. Examples of Cancer Gene Index elements with multiple child elements are given above in 1 and 5.
Special characters (e.g.for example, Parsing the Cancer Gene Index XMLMany free XML parsers exist, as do parsing modules or libraries for a variety of common programming languages, that will quickly divide the Gene-Disease and Gene-Compound XML documents into their component data. Parsed data can be stored in a database or other data management application and be computed against. Alternatively, you may prefer to write code that recursively loops through the XML and extracts the information that you desire. As end users parse the Cancer Gene Index data into various formats (e.g.for example, database dumps or tab-delimited text files) or create code to walk through the XML, they are strongly encouraged to make these versions and the code available by posting them to the Cancer Gene Index User Community Parsed Data and Code web page.
Using the Cancer Gene Index DataThe following subsections provide information and tips to maximize your use of the Cancer Gene Index XML files. Refining Your Searches with Flags and IndicatorsYou can use the Cancer Gene Index to discover associations between genes and diseases or genes and compounds. These associations were derived from the literature using a sophisticated automated process, and thus not all of the extracted gene-disease or gene-compound concept pair associations were found to be factual during validation by expert human curators in subsequent validation steps.
You also, of course, can also take advantage of other annotations . You could also filter by Codes and DetailsThe expert curators also set Evidence Codes, Role Codes, and Role Details for each concept pair. Evidence codes ( For information about the meaning of the codes, details, and other data and annotations within the XML documents, refer to the Cancer Gene Index Data, Metadata, and Annotations wiki page. Gene, Disease, and Compound OntologiesThe NCI Thesaurus provides ontological information for its concepts. Although these gene, disease, and compound (or, in NCI Thesaurus, an "agent") concept ontologies were used to construct the Cancer Gene Index Lexical Dictionaries, they are not easily deduced from information within the Cancer Gene Index, itself.
the codes, details, and other data and annotations within the XML documents, refer to the Cancer Gene Index Data, Metadata, and Annotations wiki page. Gene, Disease, and Compound OntologiesAlthough the NCI Thesaurus provides compound, disease, and gene ontological information and was used to create the data resource, this information is not easily deduced from data in the Cancer Gene Index, itself. Should you wish to search for parent, sister, and child concepts, it is possible Using the NCI Thesaurus concept terms (e.g., MatchedGeneTerm or MatchedDrugTerm) or NCI Thesaurus Concept Code (e.g. NCIDiseaseConceptCode or NCIDrugConceptCode), it is possible, however, to trace back to the hierarchical disease, compound, and gene data terms with the NCI Thesaurus graphical user interface GUI or the Enterprise Vocabulary Services (EVS) API. To use these tools, you will need the NCI Thesaurus concept terms (for example, from
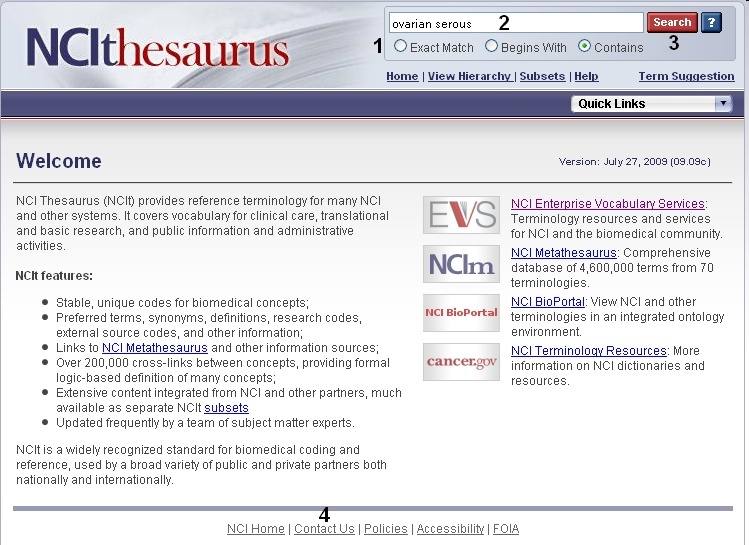
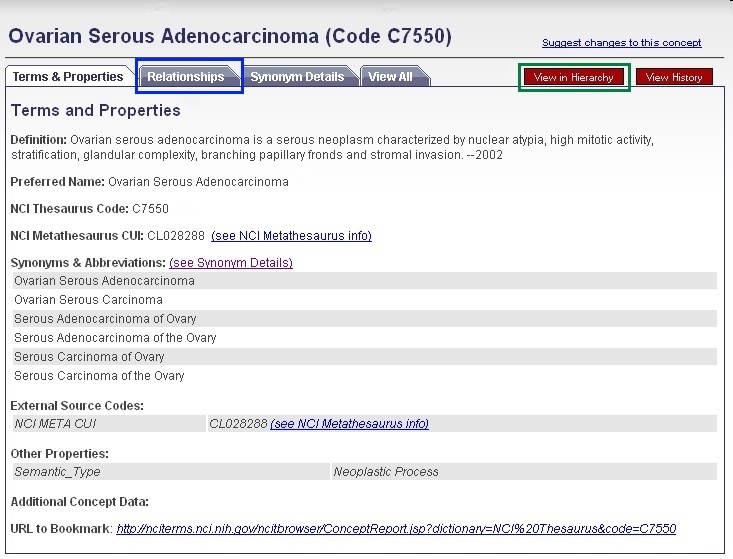
Using the NCI Thesaurus to Find Parent/Child ConceptsTo view disease, compound, and gene ontologies, open a new browser tab or window and navigate to the NCI Thesaurus web page, enter in your gene symbol, matched term, or NCI Thesaurus concept code (2, "ovarian serous adenocarcinoma"), and click the Search button (3). If you need help finding your gene, disease, or compound term, click the Contact Us link at the bottom of the page (4). You may view parent and child terms for any disease term by clicking on the Relationships tab (blue boxselected in figure). For example, "ovarian serous adenocarcinoma" has the children "ovarian serous cystadenocarcinoma" and "ovarian serous papillary adenocarcinoma" and the parent terms "malignant ovarian serous tumor," "ovarian adenocarcinoma," and "serous adenocarcinoma." Alternatively, if you would like to view where your term fits in the entire disease hierarchy, click the red View in Hierarchy button (green boxselected in figure). |