|
Page History
Adding Data to an Existing Study in caIntegrator
A Step-by-Step Illustrated Guide
This guide shows how to add clinical annotation and genomic microarray data to an existing study in caIntegrator, with a focus on common obstacles and pitfalls that may arise in the process.. It assumes that you already have basic familiarity with the program and have already created a study containing at least one source for both clinical annotation and array data. It also assumes that you have additional sources available, namely:
...
The guide is presented in a step-by-step instructional format, with each step accompanied by a screenshot from caIntegrator.
Getting Started
- Log into caIntegrator via the application's main Web page.
...
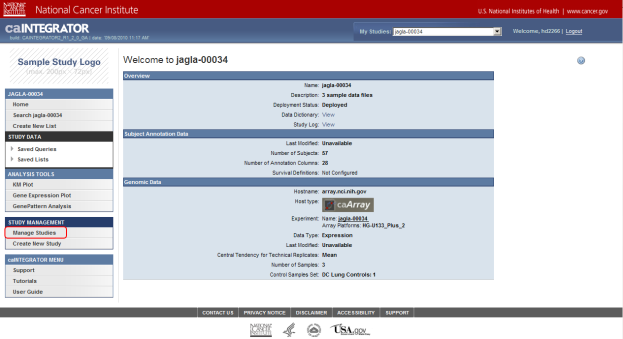
Once you log in, you are taken to the home page of the default study, which in this case is 'jagla-00034'. The study we want to add data to is 'Demo Study for ICR Folks', which you can access by clicking on the 'Manage Studies' link (highlighted in red).
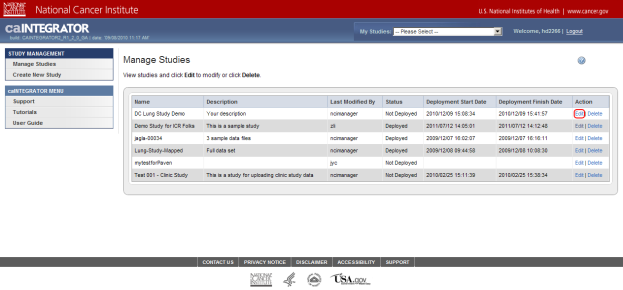
On the Manage Studies page, find the study entitled 'Demo Study for ICR Folks' in the table of studies, then click on the 'Edit' link under the Action column at the far right of the table.
You can edit the study entitled 'Demo Study for ICR Folks', which is at the top of the study list, by clicking on the 'Edit' link (highlighted in red).
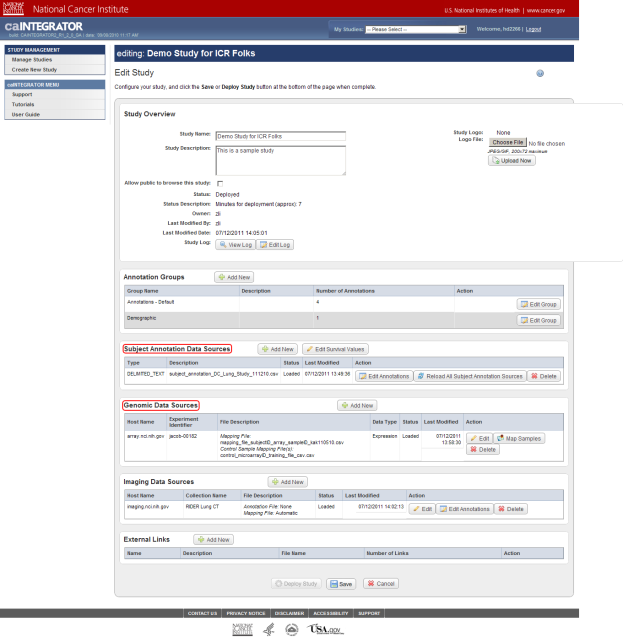
Now you are on the 'Edit Study' page, where you can modify the existing study data or load more data into it. For the purposes of this tutorial, the areas of interest on the Edit Study page are 'Subject Annotation Data Sources' and 'Genomic Data Sources', whose respective headings are highlighted in the screenshot below.
Note that this study already has some subject annotation and genomic data loaded. The annotation data is in the form of the CSV file 'subject_annotation_DC_Lung_Study_111210.CSV', while the genomic data is in the form of a link to the address of the caArray server which hosts the data (array.nci.nih.gov), as well as an experiment identifier (jacob-00182) which references the particular experiment containing the data of interest. Later in this tutorial, we will examine in depth how to load more of this data into the study.
This study already has subject annotation and genomic data loaded; they are listed beneath their respective headings, which are highlighted in red. Later in this tutorial, we'll learn how to load more data into this study.
Loading Additional Clinical Data
- Now we're ready to load additional subject annotation data into the 'Demo Study for ICR Folks'. As mentioned before, you'll need the data in the form of a CSV file containing at least one field with a unique ID for each subject in the study. The CSV file we'll use in this tutorial is called 'subject_annotations_tutorial.CSV'. A partial screenshot of the file appears below as viewed in a Microsoft Excel 2007 window.
...
This data came from a fictional multi-site study that compared gene expression between lung adenocarcinoma patients and healthy controls. The nature of the data itself is irrelevant to our purpose here. The relevant aspect is that the data is categorized into five fields, which are represented by columns in the spreadsheet.
Each field defines a different subject characteristic such as 'PATIENT_ID', which uniquely identifies each of the 100 subjects in this study (note that the screenshot above only displays data for the first 11 subjects). Once we've loaded the data into the study, we'll be able to query it by any of the fields.
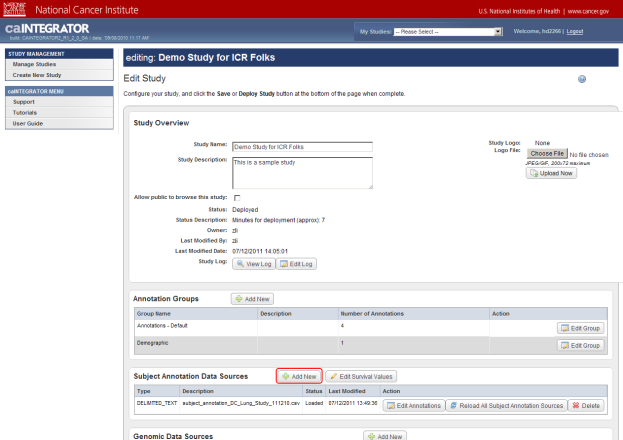
To upload your data file, first click on the 'Add New' button to the right of the 'Subject Annotation Data Sources' heading.
You can load a new subject annotation data file into the existing study by clicking on the 'Add New' button (highlighted in red).
...
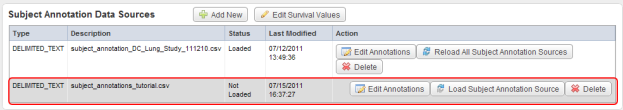
The newly uploaded source now appears in the second row (highlighted in red) of the Data Sources table. Click on the 'Load Subject Annotation Source' button under the Action column to load the source.
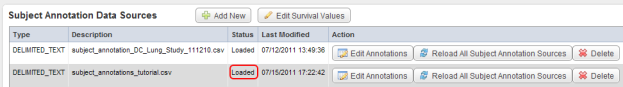
The 'Edit Study' page has now reloaded and the status of the newly added source has changed to 'Loaded' under the Status column in the Data Sources table.
The status of the newly uploaded source now appears as 'Loaded' (highlighted in red) under the Status column.
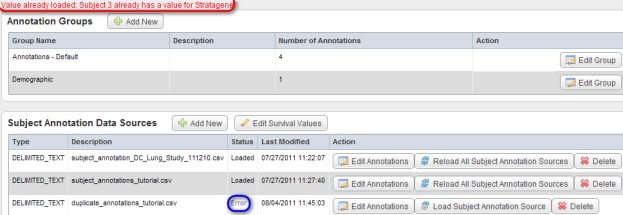
- To see what obstacles may arise in the course of loading additional data, let's try another file. This one, named 'duplicate_annotations_tutorial.CSV', contains the same five fields as each of the previously loaded files, including 'PATIENT_ID'. After repeating the procedure in steps 3 through 8, the Edit Study page displays an error message stating, "Value already loaded: Subject 3 already has a value for Stratagene" above the 'Annotation Groups' heading; in addition, the status of the newly loaded file shows as 'Error' under the 'Status' column of the 'Subject Annotation Data Sources' table.
After attempting to load the next annotation file 'duplicate_annotations_tutorial.CSV', the 'Edit Study' page shows the error message "Value already loaded: Subject 3 already has a value for Stratagene" (highlighted in red) and the status of the file shows as 'Error' (highlighted in blue).
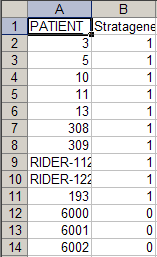
To understand why this error is occurring, let's examine the contents of the new annotation file we just tried to load. A partial screenshot of the file appears below as viewed in a Microsoft Excel 2007 window.
Notice that this file contains not only new subjects (IDs 6000 to 6002), but also some of the same subjects (i.e., IDs 3, 5, and 10) from the previously loaded file "subject_annotation_DC_Lung_Study_111210.csv". In addition, the values in the 'Stratagene' field for these subjects are different in the new file than they were in the original file. This explains the 'Value Already Loaded' error message which occurs when we attempt to load the file – this message is another way of saying that the file we're trying to load contains duplicates of subjects from previously loaded files.
We've learned a valuable lesson from this exercise: when loading additional annotation data into an existing study, make sure that your annotation file doesn't contain any duplicates of existing subjects from previously loaded files.
Querying Clinical Data
- We can't query the study unless it's already been deployed. To check whether this is the case, scroll all the way down to the bottom of the 'Edit Study' page, where you'll see a row of three buttons. If the study has been deployed, as is the case in our example, the left button labeled 'Deploy Study' will be grayed out and you will not be able to click on it. If, however, the study hasn't been deployed, the button will appear normally, and you can click on it to deploy the study.
_The bottom of the 'Edit Study' page shows the 'Deploy Study' button (highlighted in red). In this example, the study has already been deployed so this button is grayed out. If your study hasn't yet been deployed, the button will appear normally,_and you can click on it to deploy the study.
...
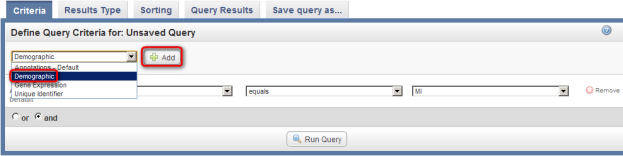
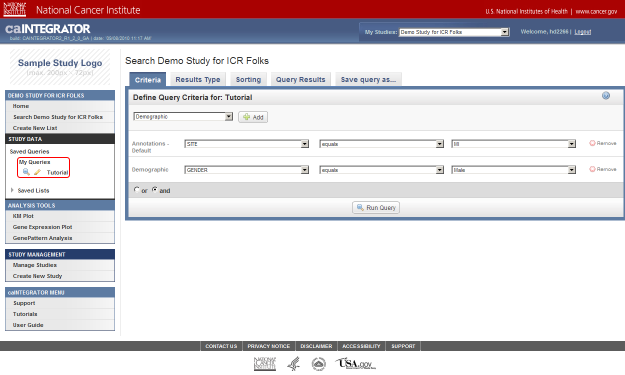
As an example, let's say we want to query the data for all male subjects located at the 'MI' study site. In this case, our two query criteria are 'Site' and 'Gender', and their respective query values are 'MI' and 'Male'. We can formulate the query by first clicking on the 'Add' button to the right of the drop-down list under the 'Define Query Criteria' heading.
To begin formulating your query, click on the 'Add' button (highlighted in red).
...
To add Gender as a field, go back to the original drop-down list (the one at the top), click on it again, click on 'Demographic' in the list, and then click on the 'Add' button to the right of the list.
Select Demographic (highlighted in red) from the drop-down list, then click on the Add button (also highlighted in red).
...
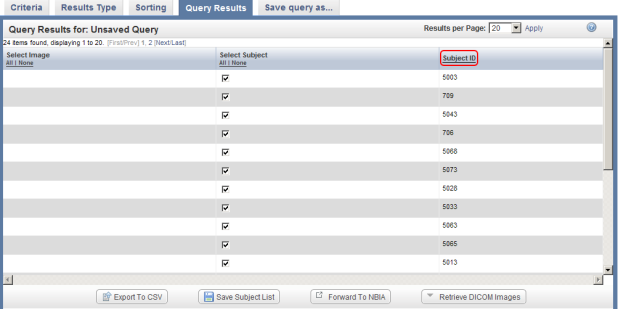
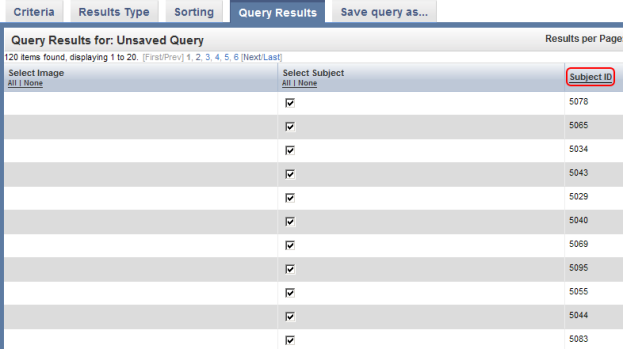
You can sort these results in numerical order of subject ID by clicking on the 'Subject ID' heading above the right table column.
You can sort query results by clicking on the Subject ID heading (highlighted in red) above the right column.
...
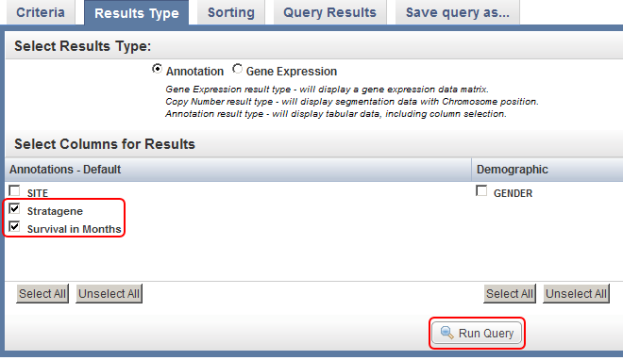
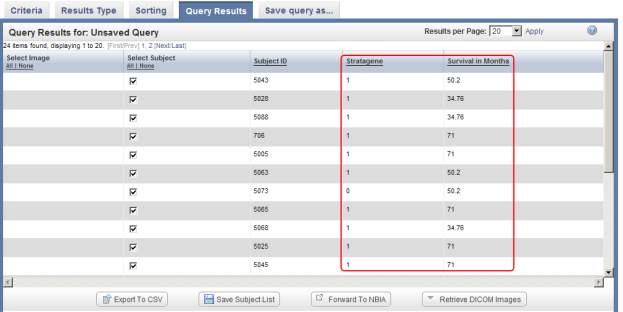
You can select additional fields (highlighted in red) to be displayed in the query results by selecting them from the checklists in the 'Results Type' tab, then clicking on the 'Run Query' button (also highlighted in red).
If you now click on the 'Run Query' button at the bottom right of the page, the results will be displayed again under the 'Query Results' tab, but this time with the additional columns Stratagene and Survival in Months, which correspond to the new fields we selected.
The updated query results include two additional columns (highlighted in red) which correspond to the two additional fields we selected in under the 'Results Type' tab.
...
The 'Tutorial' query (highlighted in red) is now saved under the 'STUDY DATA' menu in the left navigation panel and can be accessed at any time.
Loading Another Genomic Dataset
Now that you've uploaded your clinical data and learned how to query it, you're ready to do the same with your array and mapping data. To review, you'll need the server host name for your caArray data, the experiment ID, and your mapping and control training CSV files.
...
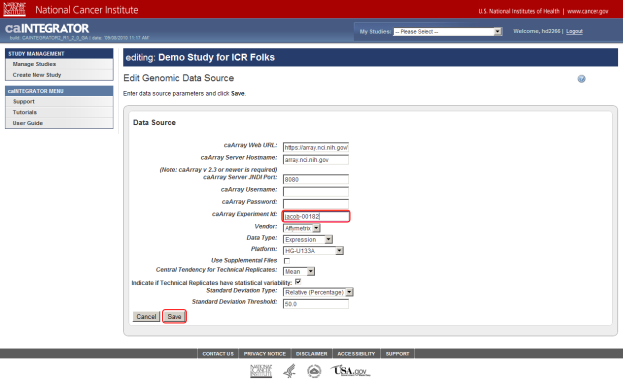
Every data source in caArray has a unique experiment ID that distinguishes it from the other sources. You can enter the ID for your experiment in the 'caArray Experiment Id' field, which is about halfway down the page. If you don't enter the ID for your source, then caIntegrator won't be able to retrieve your data, and will display an error message to that effect. In our example, the experiment ID is 'jacob-00182', which we enter in the field.
If your server hostname or any of the other values for your data source differ from the default values, then enter them into their respective fields, then click on the 'Save' button at the bottom of the page. (Remember that, if your study is private, you must enter the login credentials into the 'Username' and 'Password' fields.)
Enter the values for your data source if they differ from the default values, then click on the 'Save' button (highlighted in red). Don't forget to enter your caArray experiment ID – the ID for our example source is 'jacob-00182'.
...
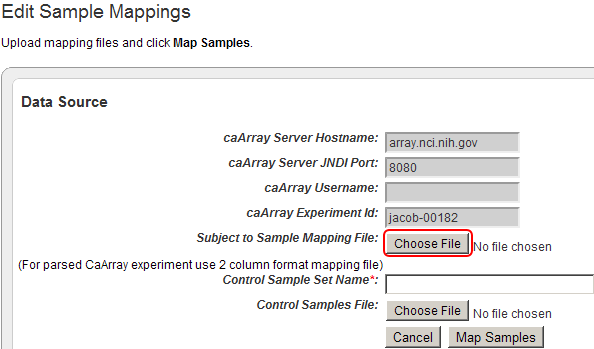
The 'Edit Sample Mappings' page shows a list of IDs for unmapped samples (highlighted in red).
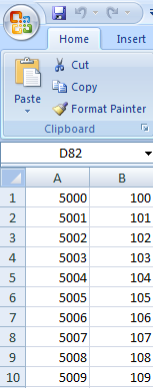
Your mapping CSV file must map the subject IDs in your annotations to the sample IDs in the unmapped samples list. A screenshot of the mapping file used in this tutorial, taken from a Microsoft Excel 2007 window, is shown below. The file is a table of two columns with no headings; the first column contains IDs of the subjects from the annotation source and the second column contains IDs from the unmapped samples list. Each subject in the left column corresponds to the sample in the right column. Note that the file doesn't map every single sample ID from the data source.
This CSV file maps the subject IDs from our annotation source (left column) to the sample IDs in our genomic source (right column).
To add your mapping CSV file to the study, click on the 'Choose File' button next to the 'Subject to Sample Mapping File' label.
Click on the 'Choose File' button (highlighted in red) to choose a mapping file to open.
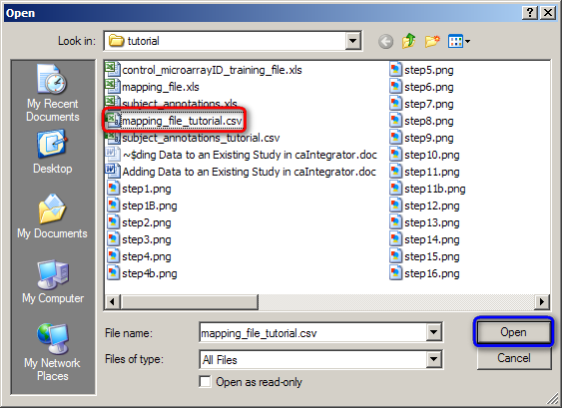
In the Open dialog that follows, find your mapping file, click on it, and then click on the 'Open' button. (In our example, the mapping file is named 'mapping_file_tutorial.CSV'.)
To open your mapping file, click on the 'mapping_file_tutorial.CSV' file (highlighted in red), then click on the 'Open' button (highlighted in blue).
...
While the 'Edit Sample Mappings' page lists all the samples from your source (both mapped and unmapped), it doesn't indicate which of these samples came from cases and which came from controls.
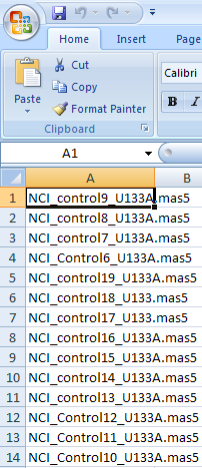
Since this information may be considered important to your study, we need a way of distinguishing between the cases and controls. The way that caIntegrator addresses this need is with a 'control training file' that lists the sample IDs of all the controls. Any sample that is not listed in this file comes from a case. The screenshot below shows a portion of an example training file in CSV format from a Microsoft Excel 2007 window.
A portion of a control training file listing the sample IDs of all the controls from our example data source. You don't need to understand the format or nomenclature of the sample IDs – they were generated by the instrument or technician who ran the samples.
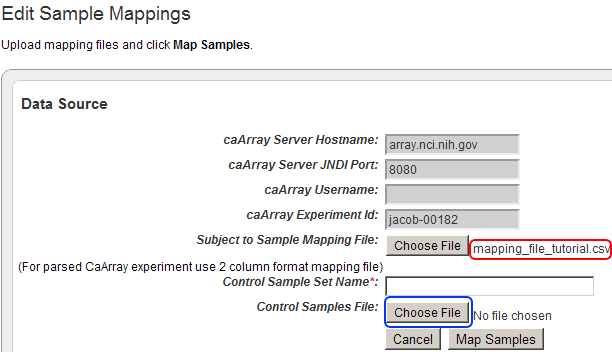
To add your control training CSV file to the study, click on the 'Choose File' button next to the 'Control Samples File' label.
The filename of the mapping file we just uploaded now appears next to the 'Choose File' button for labeled 'Subject to Sample Mapping File' (highlighted in red). Now click on the 'Choose File' button labelednext to 'Control Samples File' (highlighted in blue) to begin uploading your control training file.
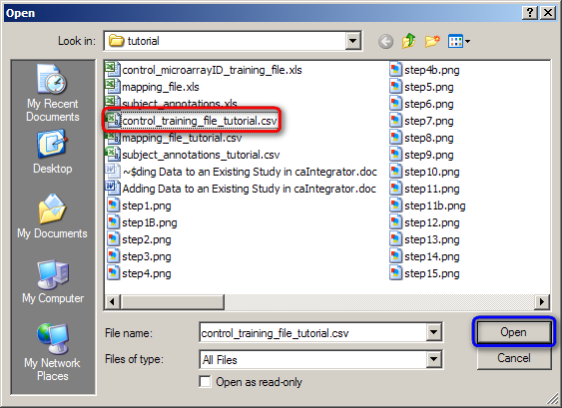
In the Open dialog that follows, find your mapping file, click on it, and then click on the 'Open' button. (In our example, the mapping file is named 'control_training_file_tutorial.CSV'.)
Click on the 'control_training_file_tutorial.CSV' file (highlighted in red), then click on the 'Open' button (highlighted in blue).
...
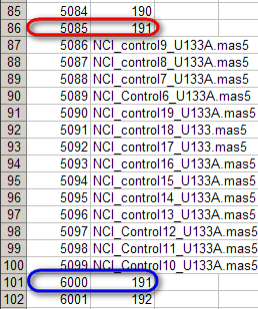
In this mapping file, the same sample (ID 191) is mapped twice, once to subject ID 5085 (highlighted in red) and again to subject ID 6000 (highlighted in blue).
You may notice something unusual about this mappings: the same sample ID (191) is mapped twice, and each mapping is to a different subject ID (5085 in one case, 6000 to another). This is obviously an error in the mappings, as each sample is taken from a single subject and must be unique to that subject. However, the question remains, what happens when we attempt to load these mappings into the study?
Suprisingly, when we repeat the procedure for loading mappings with the 'duplicate_mapping_file_tutorial.CSV', caIntegrator does not display any error message, and its source's status shows as 'Ready to be loaded' in the 'Genomic Data Sources' table, as was the case with the previous mapping file we loaded successfully. Does this mean that caIntegrator allows multiple mappings of the same sample to different subjects?
When loaded an invalid mapping file, caIntegrator does not display any error messages and shows the status of the invalidly mapped source as 'Ready to be loaded' (highlighted in red).
...
On the 'Edit Sample Mappings' page, sample ID 191 is only mapped to a single subject (highlighted in red), even though the mapping file we just loaded mapped that same sample twice.
As you can see, the mapping table shows only one mapping for sample ID 191, even though this sample was mapped to two different subjects in the new mapping file we just loaded. The subject ID it's mapped to is 6000 (the second one in the mapping file), not 5085 (the fist one in the mapping file). This means that caIntegrator overwrote the first mapping of sample ID 191 with the second one.
We've learned a valuable lesson from this exercise: be sure to check your mapping file for any duplicates before loading it into your study, as caIntegrator does not perform this check for you!
Querying Array and Mapping Data
- On the 'Edit Study' page, click on the 'My Studies' drop-down list in the blue banner at the top, then click on 'Demo Study for ICR Folks'.
...
You can sort these results in numerical order of subject ID by clicking on the 'Subject ID' heading above the right table column.
Click on the Subject ID column heading (highlighted in red) to sort the EGFR gene query results.
Click on the Subject ID column heading (highlighted in red) to sort the EGFR gene query results.
...
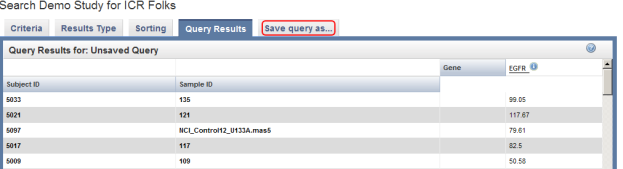
The query results now show two additional columns: Sample ID and EGFR. The latter represents median EGFR expression values. Click on the 'Save query as…' tab (highlighted in red) to save these results for future reference.
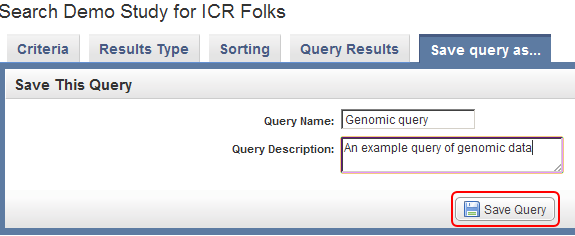
To save this query in caIntegrator for future reference, click on the 'Save query as..' tab at the top of the page, enter a name and description for the query in the respective fields, and click on the 'Save Query' button at the bottom.
Enter a query name and query description in the respective text fields, then click on the 'Save Query' button (highlighted in red) to save the query for future reference.
...