Managing Sample Composition Annotations | caNanoLab User's Guide| Managing Publications in caNanoLab
This chapter describes how to review or ascribe characterizations to samples save in caNanoLab. This section includes the following topics:
[Sample] Characterization Overview
Sample characterization, describing distinctive characteristics or essential features of the sample determined through analytical methods, records information associated with sample synthesis and properties. Samples can be characterized in caNanoLab by physical or chemical characteristics, or by data derived under in vitro and in vivo conditions.
Accessing the Sample Characterization Summary
With read-only access, you can review a summary of characterization information and annotations added to the sample.
To access characterization functions in the Navigation Tree
- Click Samples and Search Existing Samples.
- Fill in criteria, and click Search.
- Click Edit in the search results.
The Navigation Tree appears on the left sidebar and comprises functions which you can use to add annotations to the sample.
NAVIGATION TREE GENERAL INFO COMPOSITION CHARACTERIZATION PUBLICATION - General Info appears after you click the sample name and shows the Update Sample page.
- Composition defines Nanomaterial Entity, Functionalizing Entity, and Chemical Associations.
- Characterization defines essential physical characteristics that identify the material and structural properties via the Protocol and Physico-Chemical, In Vivo, and In Vitro Characterization.
- Publication displays articles, books chapters, reviews and reports already added to a sample.
Click Characterization.
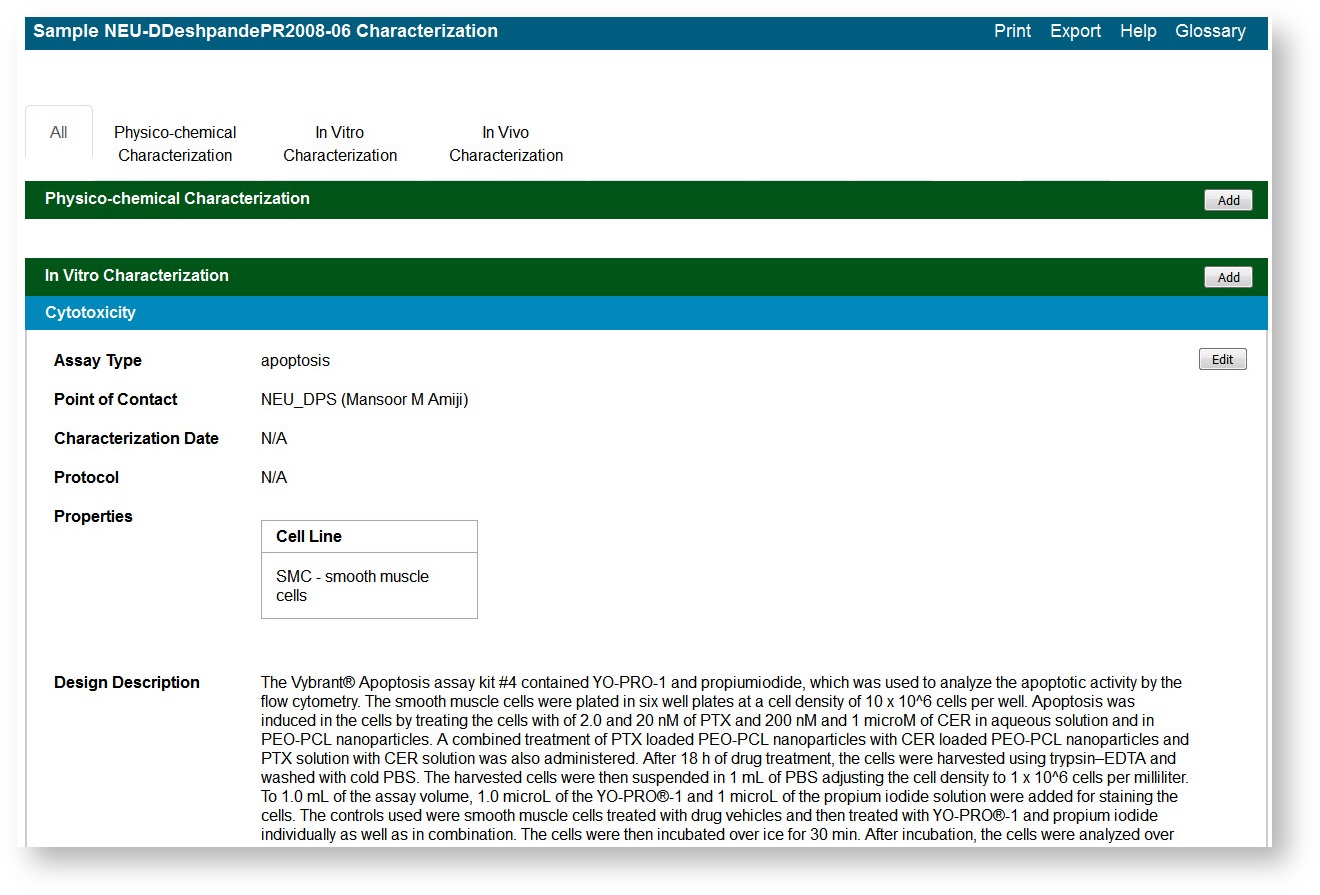
The All tab displays characterizations already added to the sample by category. Additional tabs show annotations added to the sample for each subcategory.
Submitting Characterizations
Once you access Characterizations in the Navigation Tree, you can add different types of information to the sample.
- Adding a Physico-chemical Characterization
- Adding an In Vitro Characterization
- Adding an In Vivo Characterization
If you have read-only access, you can View a Characterizations Summary.
Adding a Physico-Chemical Characterization
To add a physico-chemical characterization
- Access a sample and characterization.
Click All or the Physico-Chemical Characterizations tab. Both tabs display five sections, but All provides customizations based what you select in the Characterization Type* field.
Follow these steps to fill in the characterization. Links are provided for additional details.
Section What to Do Physico-Chemical Characterization Fill in the general information about the characterization. [Characterization] Properties Displays only for Physical State, Shape and Solubility characterizations. Define properties for the characterization, if applicable. Design and Methods Complete the fields describing techniques and instruments used to derive the data. Finding Add data findings and supporting documentation relating to the sample.
Analysis and Conclusions Enter any relevant analyses and conclusions reached by the data. Copy to other Samples... Select samples in the list to which you want this physico-chemical data transferrred. This option copies files and data to one or more selected samples "owned" by the same point of contact. When you finish, click Submit to save the data to the sample.
Defining the Physico-Chemical Characterization
Fill in the following characterization information.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
| Field | Description |
|---|---|
| Characterization Type* | The Characterization Type is already selected as Physico-Chemical Characterization. |
| Characterization Name* | Select from the drop-down list the name of the Characterization you want to add (required). |
| Assay Type | The field populates automatically with your selection in the Characterization field. |
| Protocol Name – Version | If this field is available, select from the drop-down list the protocol from which the data is derived. A hyperlink of the protocol text file may appear. Click the hyperlink to open or save the file. |
| Characterization Source | Select from the drop-down list or enter the source from which the characterization data is derived, such as a vendor or a laboratory (NCL). |
| Characterization Date | Select from the calendar or enter the date the characterization was made. Acceptable format: dd/mm/yyyy. |
Continue to define Physico-Chemical Properties.
Defining Physico-Chemical [Characterization] Properties
The [Characterization] Properties section opens only for the characterizations listed in the table below. Fill in the following Properties information as needed.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
| Characterization | Property Fields | Field Options |
|---|---|---|
Physical State | Type | Select the appropriate type from among the following options: |
Shape Properties | Type (required) | Select from the drop-down list the appropriate shape type: |
Shape Properties | Aspect Ratio | Enter the shape aspect ratio. |
Shape Properties | Minimum,/Maximum Dimensions | Enter the minimum and maximum dimensions of the sample, as well as the units of measurement. |
Solubility Properties | Solvent | Select from the options or enter the name.
|
| Solubility Properties | Critical Concentration | Enter appropriate values for the critical concentration, then select the appropriate units for those values. |
| Solubility Properties | Is Soluble? | Select Yes or No as to whether the solvent is soluble. |
Surface Properties | Is Hydrophobic? | Indicate with Yes or No whether the surface is hydrophobic. |
Continue to define Physico-Chemical Design and Methods.
Defining Physico-Chemical Design and Methods
Fill in the following Design and Methods information as needed.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
| Design and Methods Field | Description |
|---|---|
Description | Enter design and methods information not covered by other fields on the form. |
Technique and Instrument | Click Add to expand the page where you can select and enter information regarding the technique and instrument used to derive the sample. |
Technique* | Select the technique (required). |
| Abbreviation | When you select a technique, Abbreviation populates automatically if an abbreviation is known. If not, enter an abbreviation. |
| Description | Enter an appropriate description for the characterization design and methods. |
| Instrument | Click Add to expand the Instruments panel. Enter or select identifying information about the instrument used to obtain data.
|
Once you enter the data, click Save to save the information to the sample or Cancel to close the mini-window without saving data.
Continue to define Physico-Chemical Findings.
Describing Findings for Physico-Chemical Characterization
In the Finding section, you can add one or more publications relevant to the sample, as well as data derived for the sample. You can add as many files and derived data as you wish, or you can add derived data without adding files.
Click Add to expand the section, then add data and conditions and add a file.
Adding Data and Conditions
In the Data and Conditions segment of the window, you can create a matrix where you can enter data values and/or other information, such as laboratory conditions like pH or temperature that are part of your findings.
To define the data and condition matrix
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
- Add data values to Data and Conditions.
- To import a file of data values
- Save the spreadsheet of data values to a csv (comma-separated value) file.
- Click Import csv and select and follow the prompts to add the data file to the Findings Info.
- The columns and data are added to Data and Conditions.
- To add the data values manually
- Specify the number of columns and rows for the matrix, and click Update.
- Add the data values to the rows.
- Specify the number of columns and rows for the matrix, and click Update.
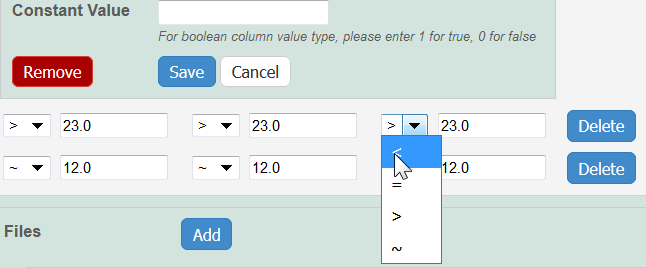
Whether you imported or added information manually, you can preface each data value with one of the following: Maintain the default, equal to (=), or select greater than (>), less than (<), or infinity (approximate).
- To import a file of data values
- To define a column, click an underlined column heading.
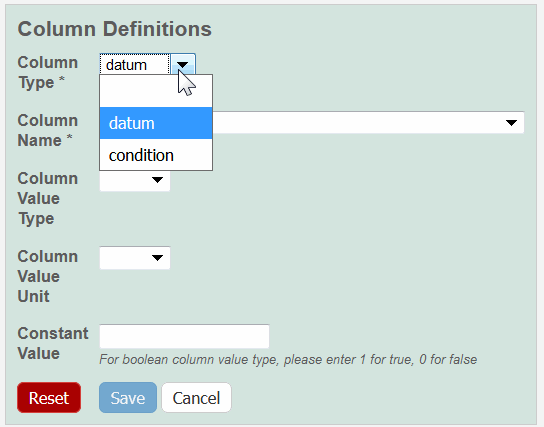
The Column Definition panel displays.
Select a Column Type, Datum or Condition.
Select a Column Name or select other and add a new one.
Column Notes
You can add up to three cell viability Column Names, including cell viability, cell viability B, and cell viability C. You can further identify the column with the Column Value Type.For Column Type, Datum, the following characterization(s) display customized Column Name options.
Characterization Type Column Type and Column Name Option(s) Physico-Chemical - Molecular – Molecular Weight
- Purity – % purity for sample
- Relaxivity – R1, R2, T1, T2
- Size – PD1, Peak N , RMS size, Z Average
- Surface – charge, zeta potential
In Vitro Enzyme Induction – % of Control
In Vivo Click Other to name the column yourself. For Column Type, Condition, all characterizations provide the Column Name options in the left column of the following table. The Column Name autopopulates the Condition Property options in the right column.
Column Type, Condition Autopopulates Column Name Column Name Autopopulates
Condition PropertyN/A
media type, serum percentage
bandwidth, frequency, time, wavelength
N/A
lyophilized, time
time
N/A
N/A
lyophilized, time
ion concentration, ionic strength, molecular formula, osmolality, serum percentage, with serum
number of pulses, pulse duration
N/A
To further identify a column, select a Column Value Type.
Once the column information is saved, the Column Type is shown in parentheses after the Column Name, such as cell viability (mean).- Select a Column Value Unit, or select other and add one.
If you want the same value to fill all rows in a column, add a Constant Value.
For Column Value Type, boolean
For Column Value Type, boolean, enter a Constant Value of 1 for true and 0 for false.Click Save, and the column(s) are updated.
If needed, click Set Column Order to change the order of the column headings in the matrix.
- Click Save in the Finding section.
Column Type "Datum" is selected with this characterization Autopopulated Column Name Option[s] Molecular Weight
Purity
Relaxivity
Size
Surface
For Column Type, Condition, all characterizations provide the Column Name options in the left column of the following table. Select a Column Name and the Condition Property options in the right column are available.
Column Type, Condition autopopulates
Column NameColumn Name autopopulates
Condition PropertyN/A
media type, serum percentage
bandwidth, frequency, time, wavelength
N/A
lyophilized, time
time
N/A
N/A
lyophilized, time
ion concentration, ionic strength, molecular formula, osmolality, serum percentage, with serum
number of pulses, pulse duration
N/A
To further identify a column, select a Column Value Type.
Once the column information is saved, the Column Type is shown in parentheses after the Column Name, such as cell viability (mean).- Column Value Unit – Select a value unit, or select other and add one.
Constant Value – If you want the same value to fill all rows in a column, add a constant value.
For Column Value Type, boolean
For Column Value Type, boolean, enter a Constant Value of 1 for true and 0 for false.
Click Save, and the column(s) are updated.
If needed, click Set Column Order to change the order of the column headings in the matrix.
- Click Save on the Finding Info panel.
After adding data and conditions to the sample, continue to add a Physico-Chemical Characterization.
Adding a File
- In the Finding section, next to Files, click Add.
- Upload, browse, and select the file or enter the file's URL where the document is located.
- Complete the following.
- Select the File Type (required), Document, Graph, Image, Movie, or Spreadsheet.
- Enter the File Title (required).
- Specify Keywords to associate with the file
- Enter a Description of additional information of the file.
Click Submit to add the file(s) to the sample.
You can add as many files as you wish.
After adding one or more files, add data and conditions or if you have no derived data to add, add a Physico-Chemical Characterization.
Adding an In Vitro Characterization
In vitro characterization allows you to add characterizations for the nanomaterial component of the sample derived from analytical techniques performed under in vitro conditions.
To add an in vitro characterization
- Access a sample and characterization.
Click All or In Vitro Characterizations. Both tabs display five sections, but the All tab provide customizations based what you select in the Characterization* field.
Follow these steps to fill in the characterization. Links are provided for additional details.
Field/Section What to Do In Vitro Characterization Fill in the general information about the characterization. [Characterization] Properties] Displays only for Cytotoxicity, Enzyme Induction and Transfection in vitro characterizations. Define properties for the characterization, if applicable. Design and Methods Complete the fields describing techniques and instruments used to derive the data. Finding Add data findings and supporting documentation relating to the sample.
Analysis and Conclusions Enter any relevant analyses and conclusions reached by the data. Copy to Other Samples... Select samples in the list to which you want this physico-chemical data transferred. This option copies files and data to one or more selected samples "owned" by the same point of contact. When you finish, click Submit to save the data to the sample.
Defining an In Vitro Characterization
Fill in the following characterization information.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
| Field | Description |
|---|---|
| Characterization Type* | The Characterization Type is already selected as In Vitro Characterization. |
| Characterization Name* | Select from the drop-down list the name of the Characterization you want to add (required). |
| Assay Type | This field populates automatically with options that display only for these in vitro characterization selections: Blood Contact, Cytotoxicity, Immune Cell Function, Oxidative Stress, Sterility and Targeting. Select an option, if appropriate, or if there are none or if you prefer, select [Other] to name the assay type. |
| Protocol Name – Version | If this field is available, select from the drop-down list the protocol from which the data is derived.
|
| Characterization Source | Select from the drop-down list or enter the source from which the characterization data is derived, such as a vendor or a laboratory (NCL). |
| Characterization Date | Select from the calendar or enter the date the characterization was made. Acceptable format: dd/mm/yyyy. |
Continue to In Vitro Properties.
Assay Type Options for In Vitro Characterizations
Specify one of the following in vitro Assay Type options or select [Other] to open a window where you can add a new assay type.
Characterization Selection | Assay Type Corresponding to Characterization |
|---|---|
Blood Contact | |
Cytotoxicity | |
Immune Cell Function | |
Oxidative stress | |
Sterility | |
Targeting | |
Transfection | Click [Other] to add an assay type. |
Return to Defining In Vitro Characterizations
Defining In Vitro [Characterization] Properties
The [Characterization] Properties section opens only when you select the following characterizations in the Summary section. To define properties for each unique characterization, enter information for the following fields.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
- For Cytotoxicity, enter the appropriate Cell Line.
- For Enzyme Induction, enter your name of choice.
- For Transfection, enter the appropriate Cell Line.
Continue to define In Vitro Design and Methods.
Defining In Vitro Design and Methods
Fill in the following Design and Methods information as needed.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
| Design and Methods Field | Description |
|---|---|
Technique and Instrument | Click Add to expand the page where you can select and enter information regarding the technique and instrument used to derive the sample. |
Technique* | Select the technique (required). |
| Abbreviation | When you select a technique, Abbreviation populates automatically if an abbreviation is known. If not, enter an abbreviation. |
| Description | Enter an appropriate description for the characterization design and methods. |
| Instrument | Click Add to expand the Instruments panel. Enter or select identifying information about the instrument used to obtain data.
|
Once you enter the data, click Save to save the information to the sample or Cancel to close the mini-window without saving data.
Continue to define In Vitro Findings.
Describing Findings for an In Vitro Characterization
In the Finding sections, you can add one or more publications relevant to the sample, as well as data derived for the sample. You can add as many files and derived data as you wish, or you can add derived data without adding files.
Click Add to expand the section, then add data and conditions and add a file.
Adding Data and Conditions
In the Data and Conditions segment of the window, you can create a matrix where you can enter data values and/or other information, such as laboratory conditions that are part of your findings.
To define the matrix for the data, follow these steps.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
- Add data values to Data and Conditions.
- To import a file of data values
- Save the spreadsheet of data values to a csv (comma-separated value) file.
- Click Import csv and select and follow the prompts to add the data file to the Findings Info.
- The columns and data are added to Data and Conditions.
- To add the data values manually
- Specify the number of columns and rows for the matrix, and click Update.
- Add the data values to the rows.
- Specify the number of columns and rows for the matrix, and click Update.
Whether you imported or added information manually, you can preface each data value with one of the following: Maintain the default, equal to (=), or select greater than (>), less than (<), or infinity (approximate).
- To import a file of data values
- To define a column, click an underlined column heading.
The Column Definition panel displays.
Select a Column Type, Datum or Condition.
Select a Column Name or select other and add a new one.
Column Notes
You can add up to three cell viability Column Names, including cell viability, cell viability B, and cell viability C. You can further identify the column with the Column Value Type.For Column Type, Datum, the following characterization(s) display customized Column Name options.
Characterization Type Column Type and Column Name Option(s) Physico-Chemical - Molecular – Molecular Weight
- Purity – % purity for sample
- Relaxivity – R1, R2, T1, T2
- Size – PD1, Peak N , RMS size, Z Average
- Surface – charge, zeta potential
In Vitro Enzyme Induction – % of Control
In Vivo Click Other to name the column yourself. For Column Type, Condition, all characterizations provide the Column Name options in the left column of the following table. The Column Name autopopulates the Condition Property options in the right column.
Column Type, Condition Autopopulates Column Name Column Name Autopopulates
Condition PropertyN/A
media type, serum percentage
bandwidth, frequency, time, wavelength
N/A
lyophilized, time
time
N/A
N/A
lyophilized, time
ion concentration, ionic strength, molecular formula, osmolality, serum percentage, with serum
number of pulses, pulse duration
N/A
To further identify a column, select a Column Value Type.
Once the column information is saved, the Column Type is shown in parentheses after the Column Name, such as cell viability (mean).- Select a Column Value Unit, or select other and add one.
If you want the same value to fill all rows in a column, add a Constant Value.
For Column Value Type, boolean
For Column Value Type, boolean, enter a Constant Value of 1 for true and 0 for false.Click Save, and the column(s) are updated.
If needed, click Set Column Order to change the order of the column headings in the matrix.
- Click Save in the Finding section.
Column Type "Datum" is selected with this characterization Autopopulated Column Name Option[s] Enzyme Induction
For Column Type, Condition, all characterizations provide the Column Name options in the left column of the following table. Select a Column Name and the Condition Property options in the right column are available.
Column Type, Condition autopopulates
Column NameColumn Name autopopulates
Condition PropertyN/A
media type, serum percentage
bandwidth, frequency, time, wavelength
N/A
lyophilized, time
time
N/A
N/A
lyophilized, time
ion concentration, ionic strength, molecular formula, osmolality, serum percentage, with serum
number of pulses, pulse duration
N/A
To further identify a column, select a Column Value Type.
Once the column information is saved, the Column Type is shown in parentheses after the Column Name, such as cell viability (mean).- Column Value Unit – Select a value unit, or select other and add one.
Constant Value – If you want the same value to fill all rows in a column, add a constant value.
For Column Value Type, boolean
For Column Value Type, boolean, enter a Constant Value of 1 for true and 0 for false.
Click Save, and the column(s) are updated.
If needed, click Set Column Order to change the order of the column headings in the matrix.
- Click Save on the Finding Info panel.
After adding data and conditions to the sample, add an In Vitro Characterization.
Adding a File
- In the Finding section, next to Files, click Add.
- Upload, browse, and select the file or enter the file's URL where the document is located.
- Complete the following.
- Select the File Type (required), Document, Graph, Image, Movie, or Spreadsheet.
- Enter the File Title (required).
- Specify Keywords to associate with the file
- Enter a Description of additional information of the file.
Click Submit to add the file(s) to the sample.
You can add as many files as you wish.
Adding an In Vivo Characterization
In vivo characterization allows you to add characterizations for the nanomaterial component of the sample that were derived from analytical techniques performed under in vivo conditions.
To add an in vivo characterization
- Access a sample and characterization.
Click the All or In Vivo Characterizations tab. Both tabs display five sections, but the All tab provide customizations based what you select in the Characterization* field.
Follow these steps to fill in the characterization. Links are provided for additional details.
Field/Section What to Do In Vivo Characterization Fill in the general information about the characterization. [Characterization] Properties Displays for in vivo characterizations. Design and Methods Complete the fields describing techniques and instruments used to derive the data. Findings Add data findings and supporting documentation relating to the sample.
Analysis and Conclusions Enter any relevant analyses and conclusions reached by the data. Copy to Other Samples... Select samples in the list to which you want this physico-chemical data transferred. This option copies files and data to one or more selected samples "owned" by the same point of contact.
When you finish, click Submit to save the data to the sample.
Defining an In Vivo Characterization
Fill in the following characterization information.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
| Field | Description |
|---|---|
| Characterization Type* | The Characterization Type is already selected as In Vivo Characterization. |
| Characterization Name* | Select from the drop-down list the name of the Characterization you want to ad (required). The options for In Vivo Characterization are Pharmacokinetics and Toxicology. There are no customizations on this Characterization page based on either of these selections. |
| Assay Type | Leave this blank or select a type. |
| Protocol Name – Version | Select from the drop-down list the protocol from which the data is derived. A hyperlink of the protocol text file may appear. Click the hyperlink to open or save the file. |
| Characterization Source | Select from the drop-down list or enter the source from which the characterization data is derived, such as a vendor or a laboratory (NCL). |
| Characterization Date | Select from the calendar or enter the date the characterization was made. Acceptable format: dd/mm/yyyy. |
Continue to define In Vivo Design and Methods.
Defining In Vivo Design and Methods
Fill in the following Design and Methods information as needed.
To enter an alternative to an option, select other if available and enter a value. The value is added to the list of options.
| Design and Methods Field | Description |
|---|---|
Description | Enter information not covered by other fields on the form. |
Technique and Instrument | Click Add to expand the page where you can select and enter information regarding the technique and instrument used to derive the sample. |
Technique* | Select the technique (required). |
| Abbreviation | When you select a technique, Abbreviation populates automatically if an abbreviation is known. If not, enter an abbreviation. |
| Description | Enter an appropropriate description for the design and methods. |
| Instrument | Click Add to expand the Instruments panel. Enter or select identifying information about the instrument used to obtain data.
|
Once you specify the data, click Save to save the information to the sample or Cancel to close the mini-window without saving data.
Continue to define In Vivo Findings.
Describing Findings for In Vivo Characterizations
In the Finding sections, you can add one or more publications relevant to the sample as well as data derived for the sample. You can add as many files and derived data as you wish, or you can add derived data without adding files.
Complete the instructions as described in Adding Data and Conditions and Adding a File.
Adding Data and Conditions
In the Data and Conditions segment of the window, you can create a matrix where you can enter data values and/or other information that are part of your findings.
To define the matrix for the data
- Add data values to Data and Conditions.
- To import a file of data values
- Save the spreadsheet of data values to a csv (comma-separated value) file.
- Click Import csv and select and follow the prompts to add the data file to the Findings Info.
- The columns and data are added to Data and Conditions.
- To add the data values manually
- Specify the number of columns and rows for the matrix, and click Update.
- Add the data values to the rows.
- Specify the number of columns and rows for the matrix, and click Update.
Whether you imported or added information manually, you can preface each data value with one of the following: Maintain the default, equal to (=), or select greater than (>), less than (<), or infinity (approximate).
- To import a file of data values
- To define a column, click an underlined column heading.
The Column Definition panel displays.
Select a Column Type, Datum or Condition.
Select a Column Name or select other and add a new one.
Column Notes
You can add up to three cell viability Column Names, including cell viability, cell viability B, and cell viability C. You can further identify the column with the Column Value Type.For Column Type, Datum, the following characterization(s) display customized Column Name options.
Characterization Type Column Type and Column Name Option(s) Physico-Chemical - Molecular – Molecular Weight
- Purity – % purity for sample
- Relaxivity – R1, R2, T1, T2
- Size – PD1, Peak N , RMS size, Z Average
- Surface – charge, zeta potential
In Vitro Enzyme Induction – % of Control
In Vivo Click Other to name the column yourself. For Column Type, Condition, all characterizations provide the Column Name options in the left column of the following table. The Column Name autopopulates the Condition Property options in the right column.
Column Type, Condition Autopopulates Column Name Column Name Autopopulates
Condition PropertyN/A
media type, serum percentage
bandwidth, frequency, time, wavelength
N/A
lyophilized, time
time
N/A
N/A
lyophilized, time
ion concentration, ionic strength, molecular formula, osmolality, serum percentage, with serum
number of pulses, pulse duration
N/A
To further identify a column, select a Column Value Type.
Once the column information is saved, the Column Type is shown in parentheses after the Column Name, such as cell viability (mean).- Select a Column Value Unit, or select other and add one.
If you want the same value to fill all rows in a column, add a Constant Value.
For Column Value Type, boolean
For Column Value Type, boolean, enter a Constant Value of 1 for true and 0 for false.Click Save, and the column(s) are updated.
If needed, click Set Column Order to change the order of the column headings in the matrix.
- Click Save in the Finding section.
For an in vivo characterization, click [Other] to name the column yourself.
For Column Type, Condition, all characterizations provide the Column Name options in the left column of the following table. Select a Column Name and the Condition Property options in the right column are available.
Column Type, Condition autopopulates
Column NameColumn Name autopopulates
Condition PropertyN/A
media type, serum percentage
bandwidth, frequency, time, wavelength
N/A
lyophilized, time
time
N/A
N/A
lyophilized, time
ion concentration, ionic strength, molecular formula, osmolality, serum percentage, with serum
number of pulses, pulse duration
N/A
To further identify a column, select a Column Value Type.
Once the column information is saved, the Column Type is shown in parentheses after the Column Name, such as cell viability (mean).- Column Value Unit – Select a value unit, or select other and add one.
Constant Value – If you want the same value to fill all rows in a column, add a constant value.
For Column Value Type, boolean
For Column Value Type, boolean, enter a Constant Value of 1 for true and 0 for false.
Click Save, and the column(s) are updated.
If needed, click Set Column Order to change the order of the column headings in the matrix.
- Click Save on the Finding Info panel.
Adding a File
- In the Finding section, next to Files, click Add.
- Upload, browse, and select the file or enter the file's URL where the document is located.
- Complete the following.
- Select the File Type (required), Document, Graph, Image, Movie, or Spreadsheet.
- Enter the File Title (required).
- Specify Keywords to associate with the file
- Enter a Description of additional information of the file.
Click Submit to add the file(s) to the sample.
You can add as many files as you wish.
After adding one or more files, continue by Adding Data and Conditions or if you have no derived data to add, return to the steps described in Adding an In Vivo Characterization.
Characterization Tasks
From a characterization summary page, and with curator privileges, you can perform these tasks:
- Printing a Characterization
- Exporting Characterizations
- Editing Characterizations
- Copying Characterizations
- Deleting a Characterization
Printing a Characterization
To print a characterization
- Access a sample and characterization.
- The All tab displays the characterizations summary.
Click Print at the top right of the page.
Read only access?
If buttons described in this section do not display, then you can assume that you have read-only access to the data.
Exporting Characterizations
To export a characterization
- Access a sample and characterization.
- The All tab displays the characterizations summary.
Click Export at the top right of the page.
Read only access?
If buttons described in this section do not display, then you can assume that you have read-only access to the data.
Editing Characterizations
To edit a characterization
- Access a sample and characterization.
- The All tab displays the characterizations summary.
Click Edit at the right of the characterization section you want to change. This opens the corresponding Characterization page where you can edit the file by following the same directions as described for creating characterizations.
Read only access?
If buttons described in this section do not display, then you can assume that you have read-only access to the data.
Copying Characterizations
The characterization files and/or derived data for this sample can be copied to other samples from the same primary point of contact.
To copy characterizations, open the sample characterization form that displays the characterization
- Access a sample and characterization.
- The All tab displays the characterizations summary.
- On the characterization summary page, click Edit at the right of the section you want to copy. This opens the corresponding Characterization page.
- At the bottom of the page, in the Copy to other samples... box, select one or more samples that have the same primary point of contact (those listed).
- Click Also copy characterization results? .
Click Submit to copy the characterization.
Read only access?
If buttons described in this section do not display, then you can assume that you have read-only access to the data.
Deleting a Characterization
To delete a characterization for a selected sample
Deleting a characterization
This only deletes the characterization(s) for the selected sample. If the characterization(s) is copied for other samples, the characterization will not be deleted.
- Access a sample and characterization.
- The All tab displays the characterizations summary.
- Click Edit at the right of the section you want to delete. This opens the corresponding Characterization page.
Click Delete at the bottom left of the page.
Read only access?
If buttons described in this section do not display, then you can assume that you have read-only access to the data.
Deleted characterizations are placed in the sample archive for history purposes.